Experiment
Last compiled on 23-03-2026
1 Getting started
To copy the code, click the button in the upper right corner of the code-chunks.
1.1 clean up
rm(list = ls())
gc()1.2 custom functions
We defined a number custom functions, at Download custom_functions.R.
source("./custom_functions.R")1.3 necessary packages
tidyverse: data wranglingigraph: generate and visualize graphsparallel: parallel computing to speed up simulationforeach: looping in paralleldoParallel: parallel backend forforeachggplot2: data visualizationggh4x: hacks forggplot2ggpubr: make visualizations publication-ready
packages = c("tidyverse", "igraph", "ggplot2", "parallel", "doParallel", "foreach", "ggh4x", "ggpubr",
"plotly", "RColorBrewer", "grid", "gridExtra", "patchwork", "ggplotify", "ggraph", "gganimate", "RColorBrewer",
"ggtext", "magick", "jsonlite")
invisible(fpackage.check(packages))
rm(packages)2 Experimental conditions
2.1 1. heavy-tailed network with central minorities
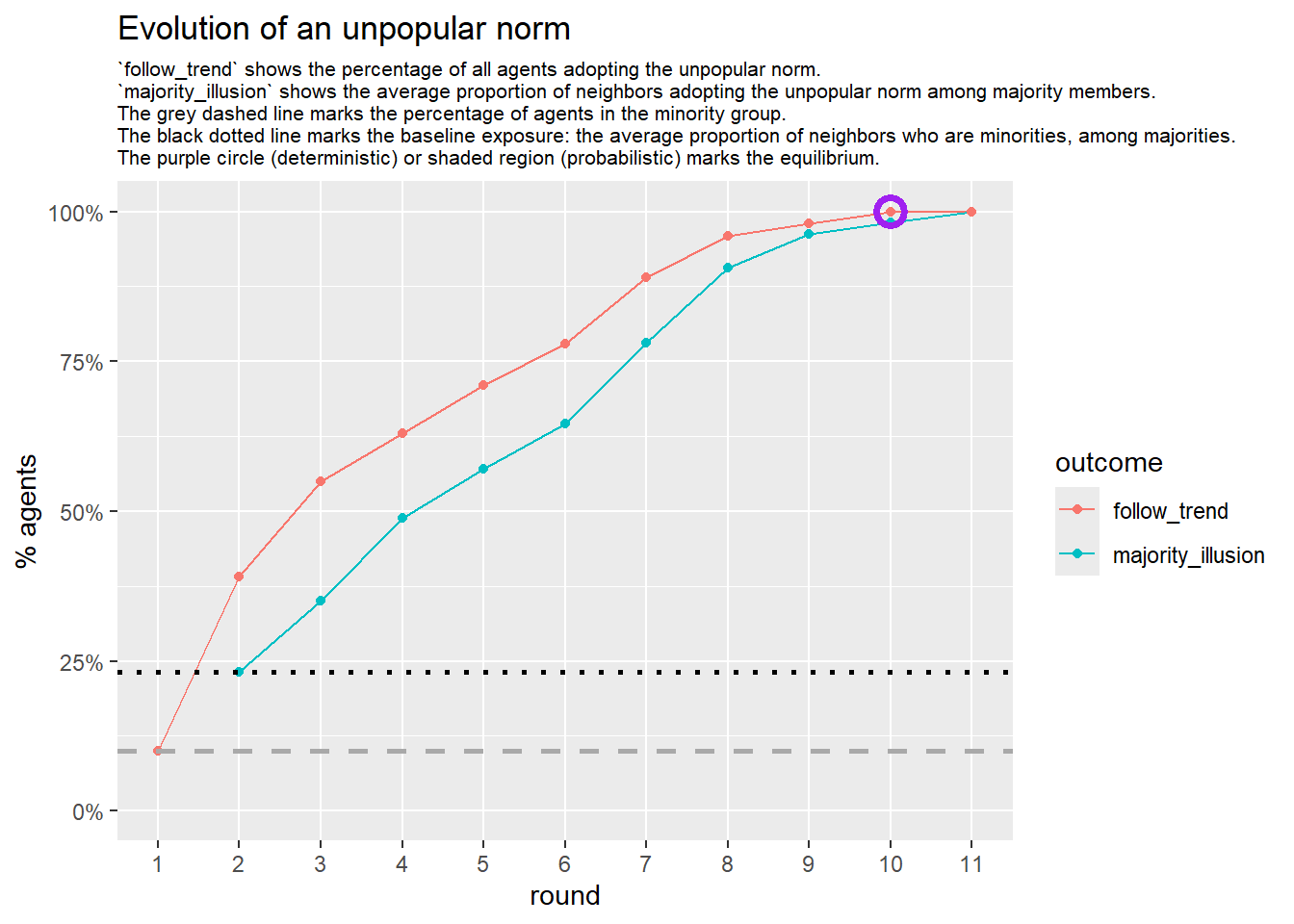
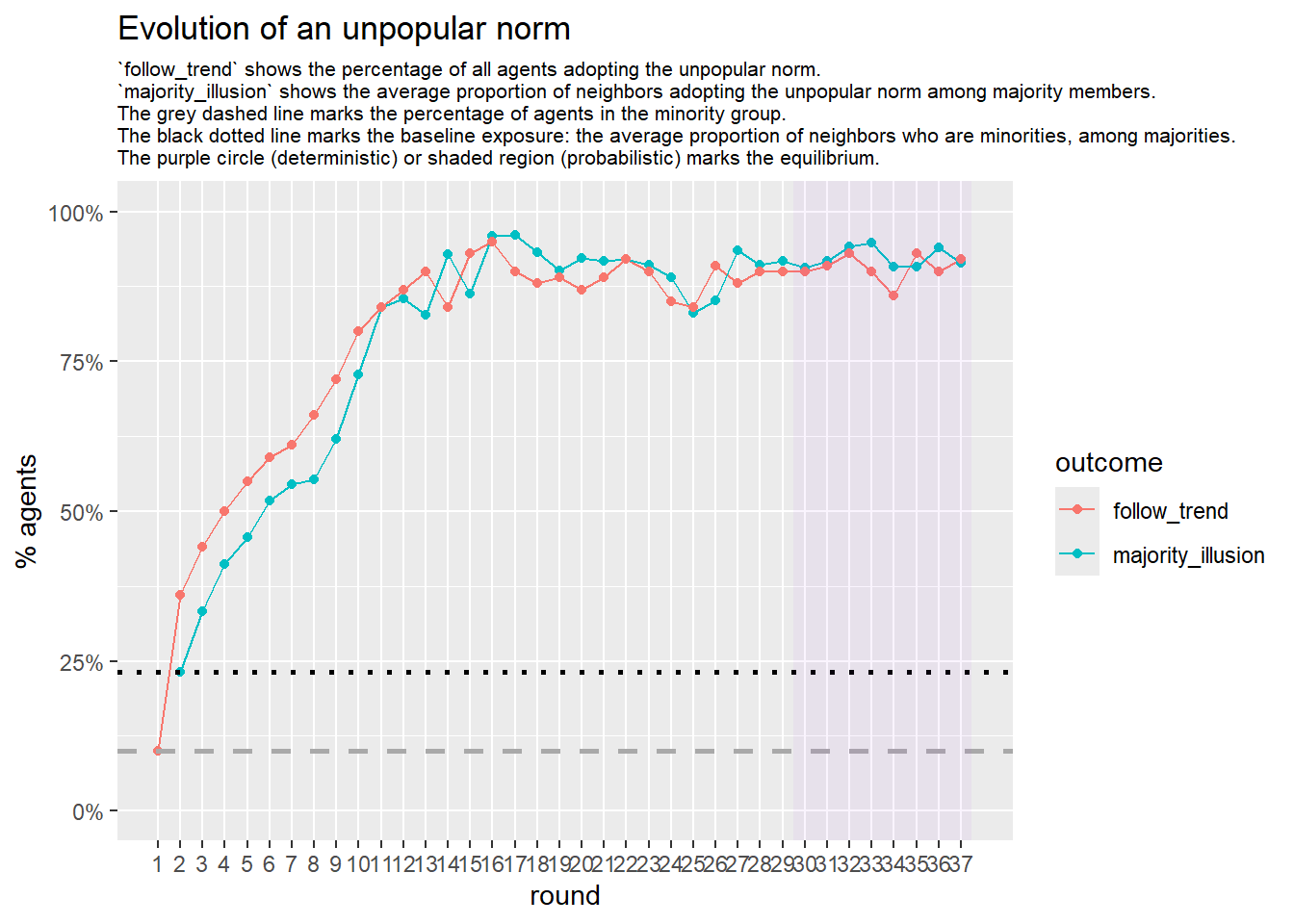
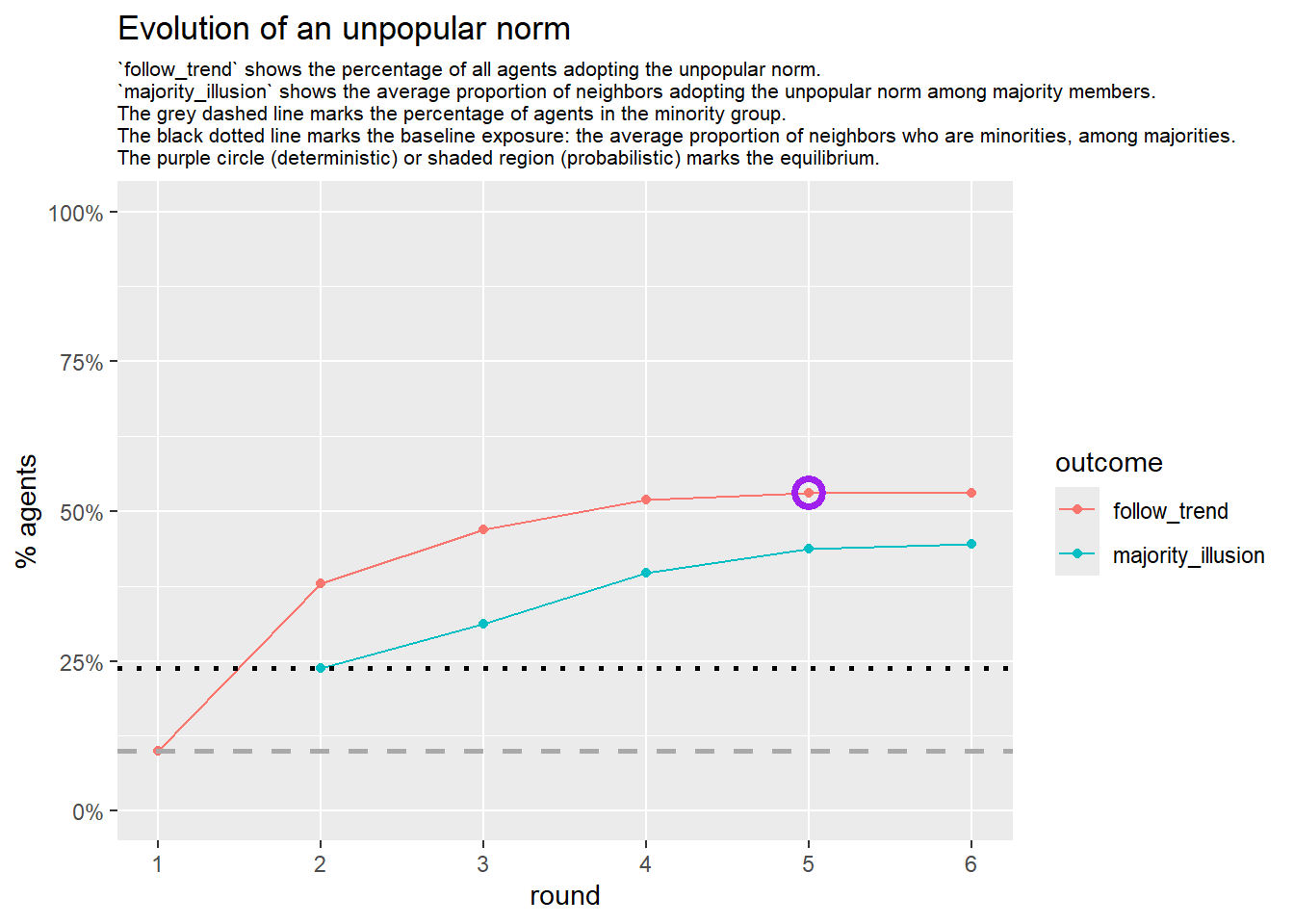
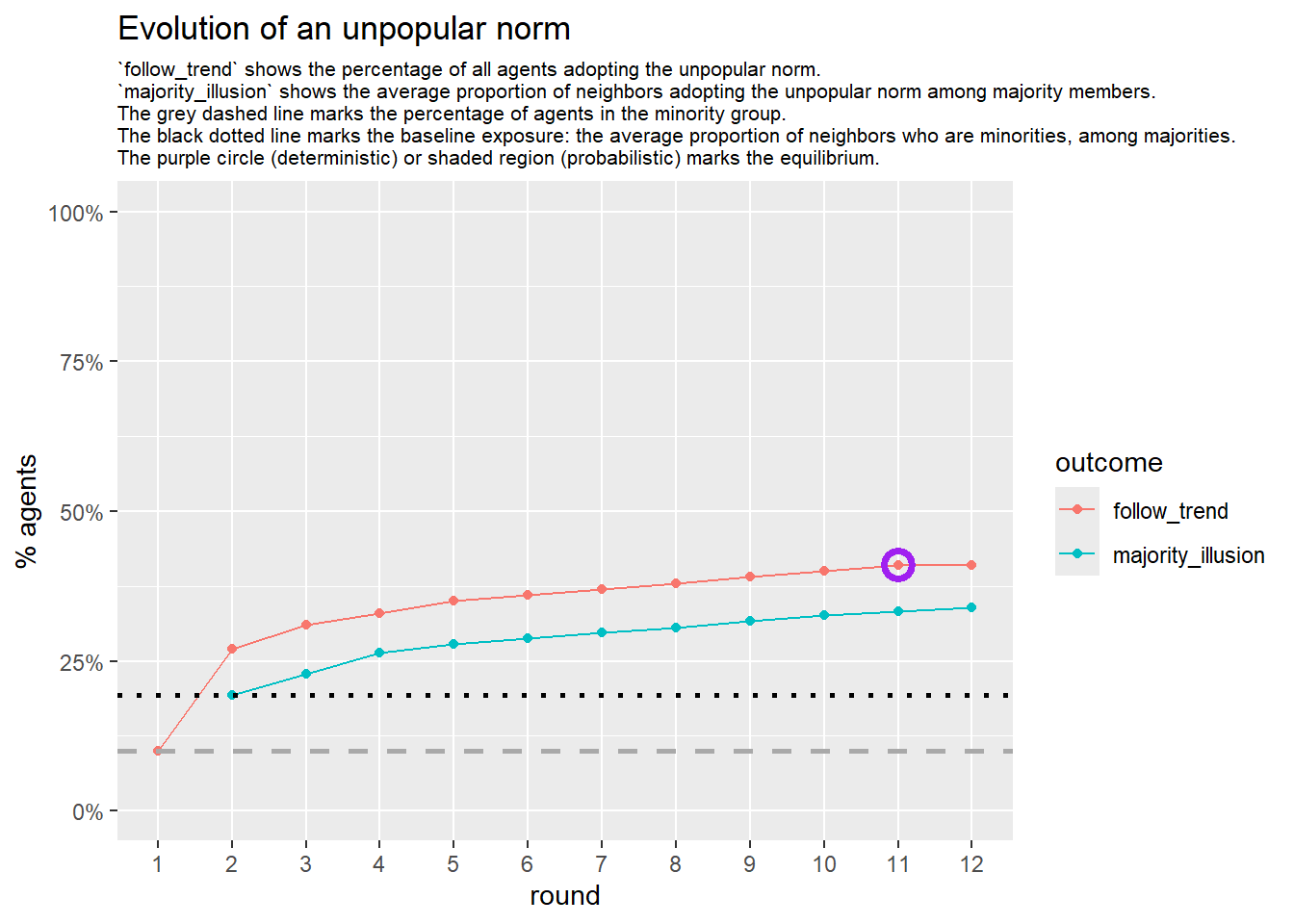
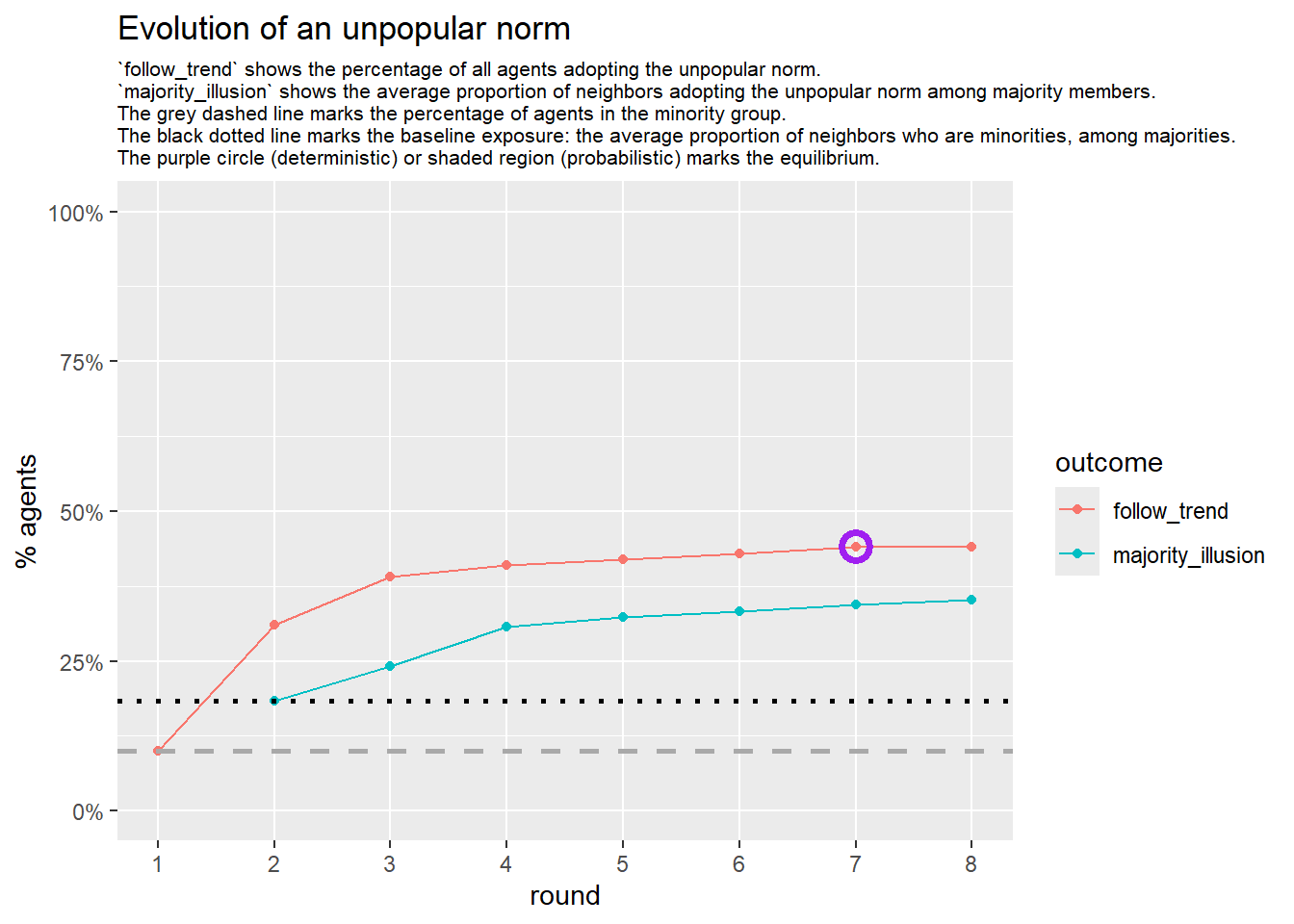
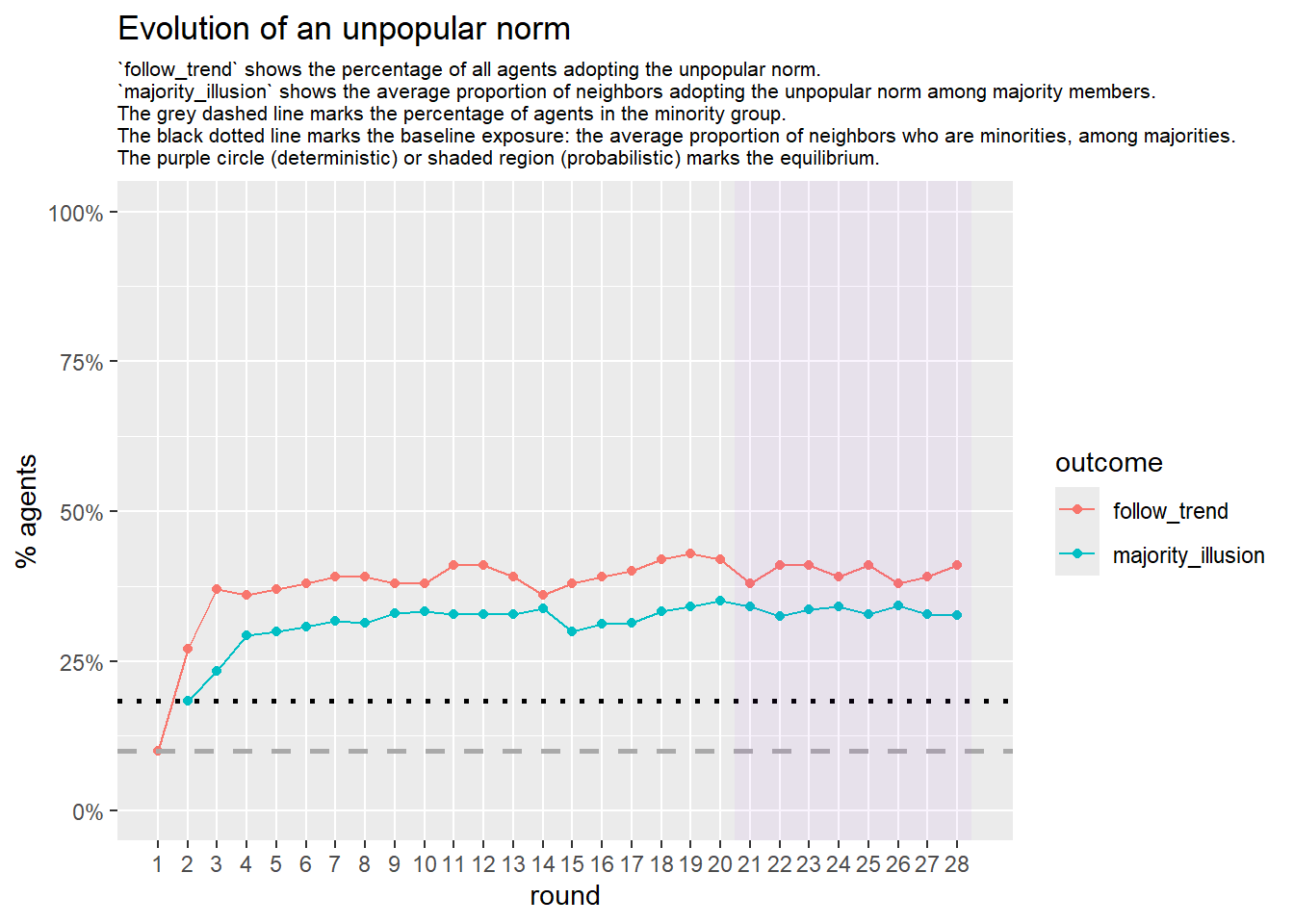
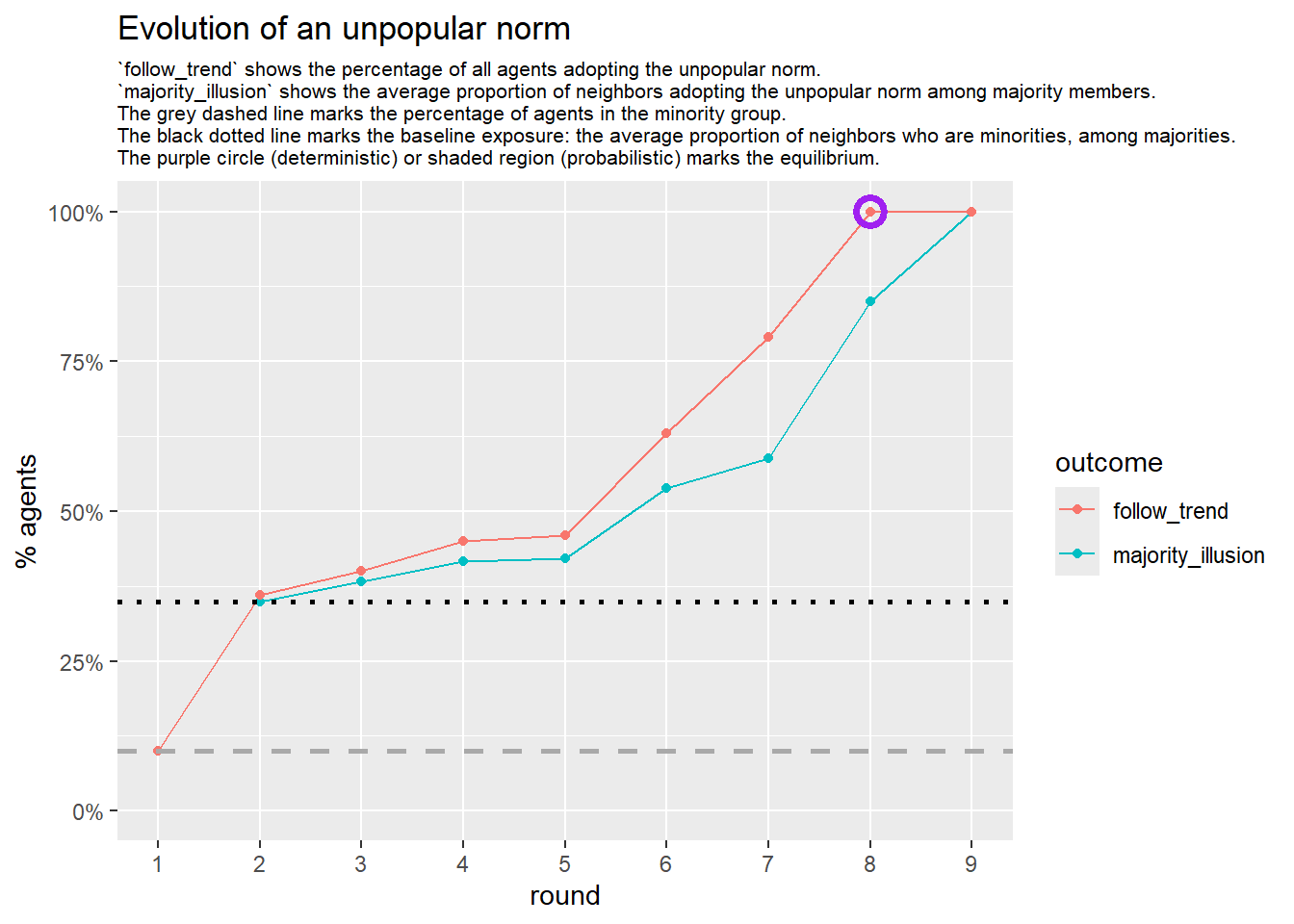
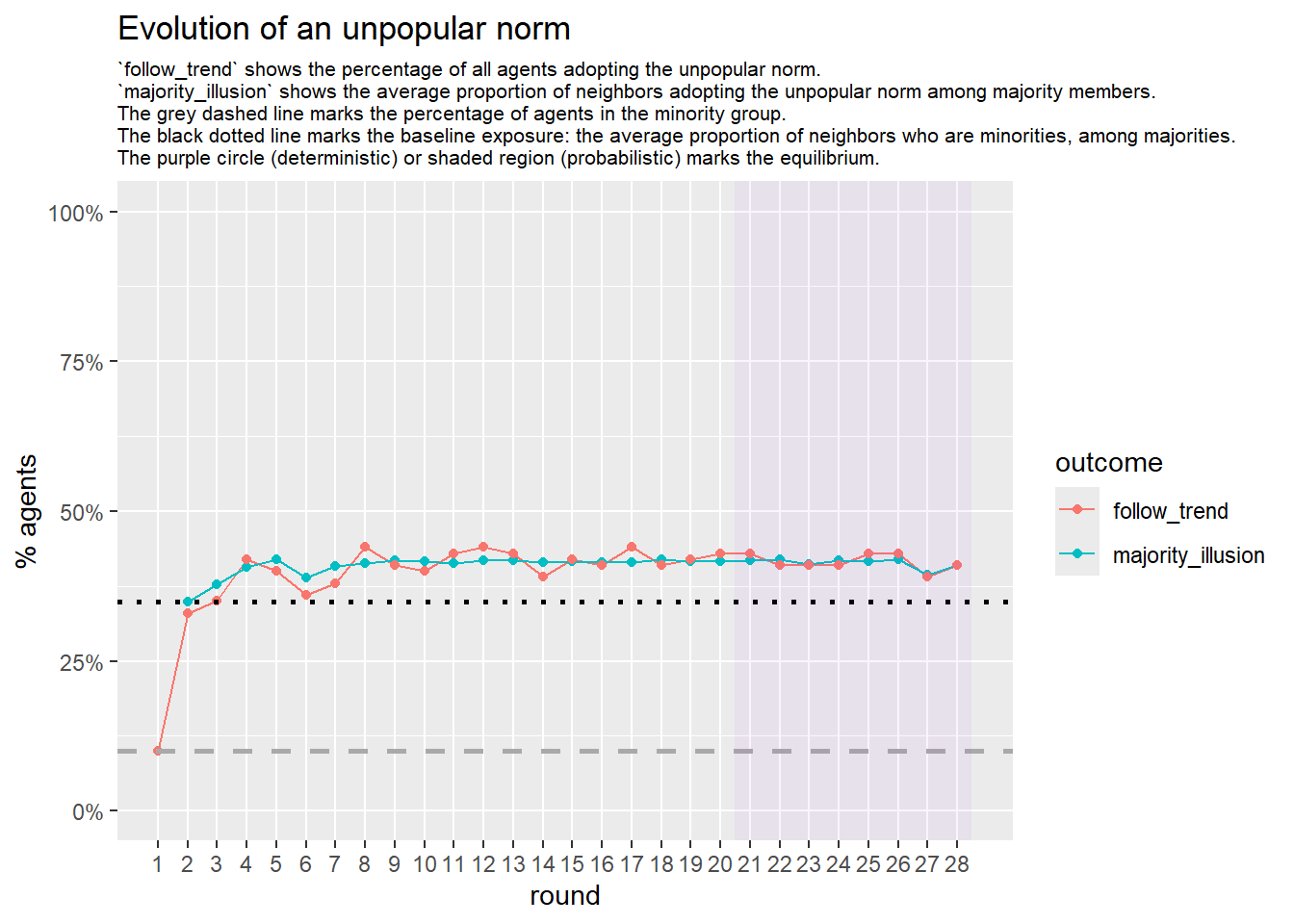
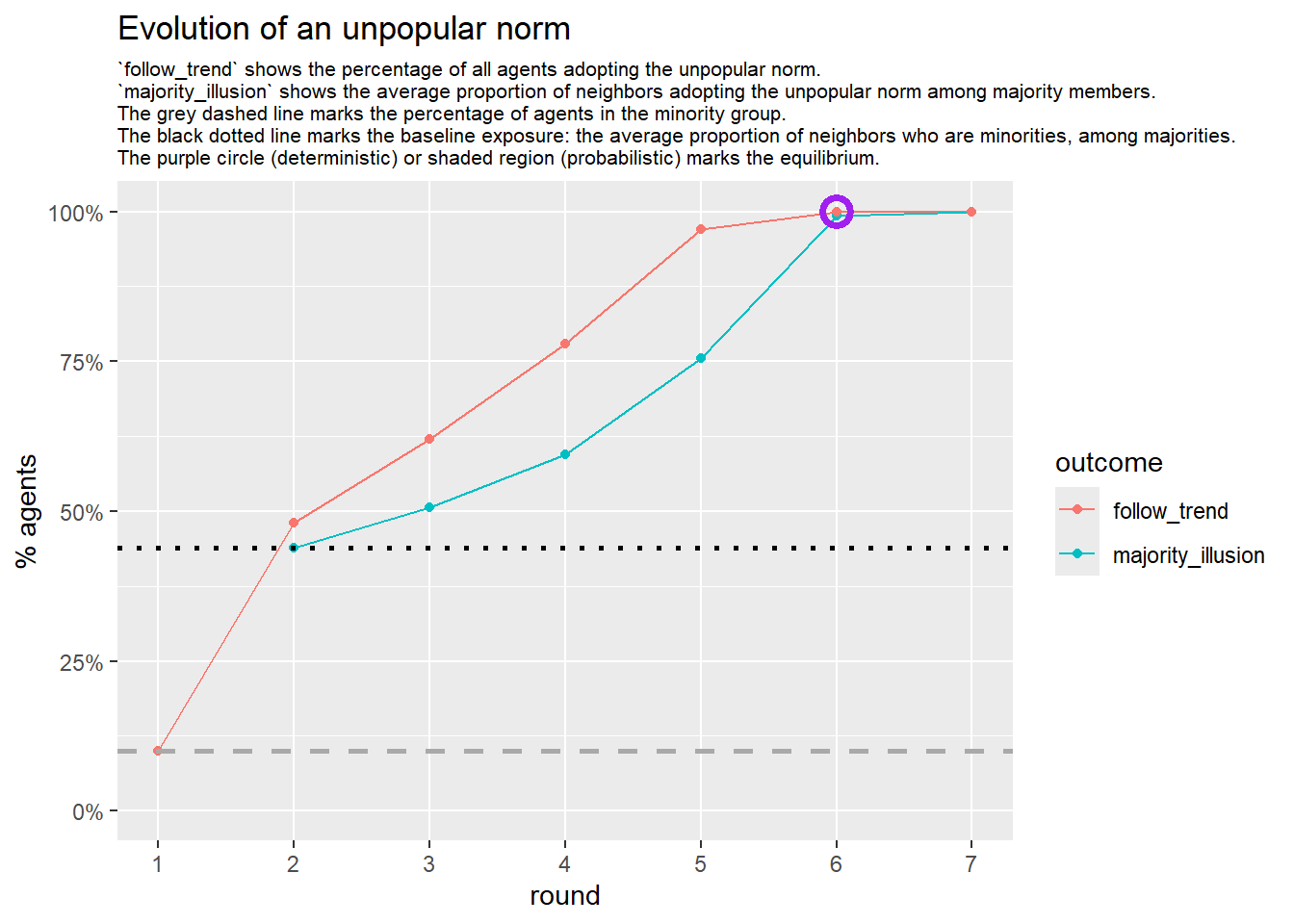
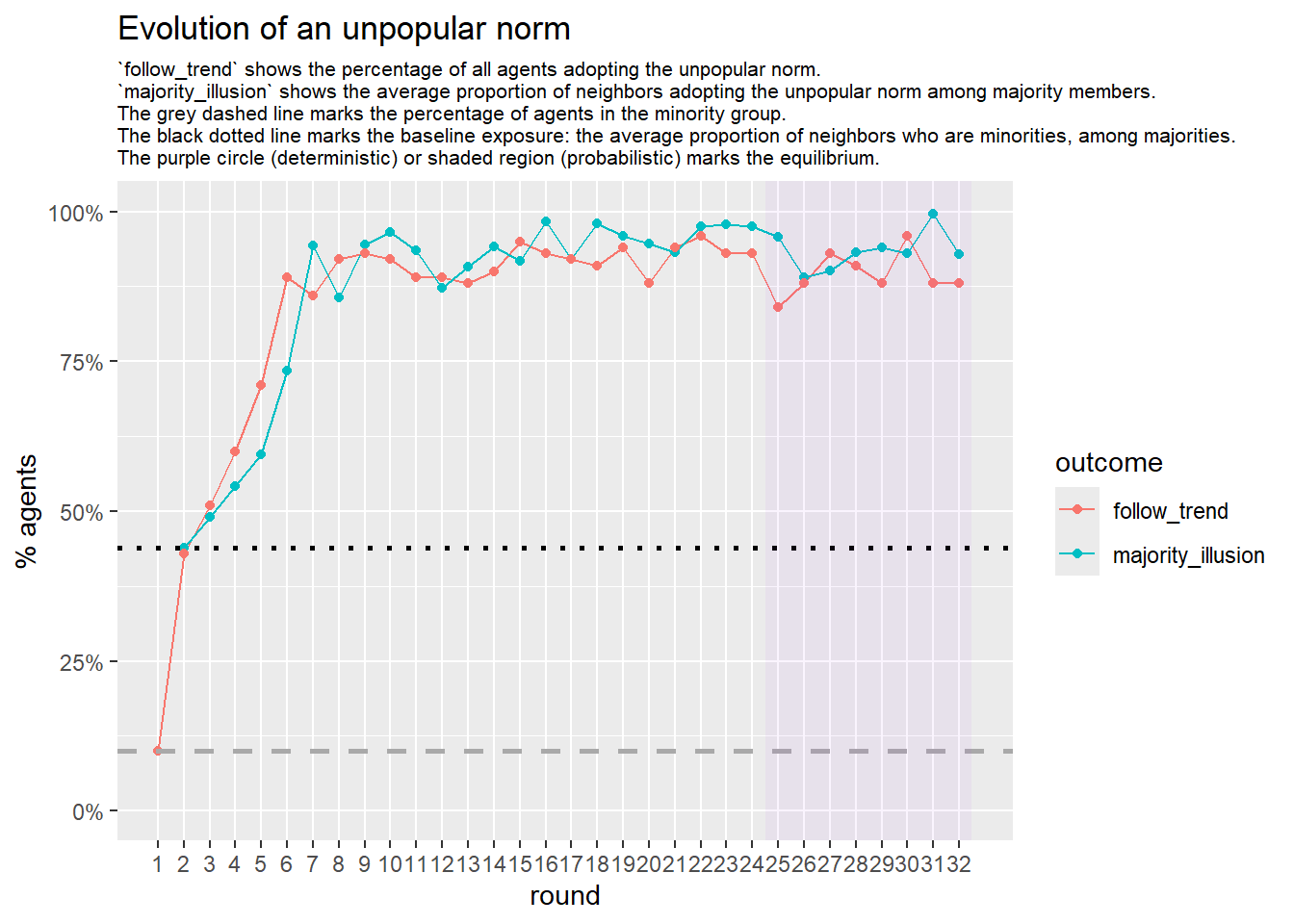
Let’s take a likely-case for an unpopular norm, generate multiple networks of this target, and simulate the emergence of the norm following a deterministic and probabilistic choice model:
# pick one configuration that likely leads to an unpopular norm, and explore multiple 'seeds':
run_one_seed <- function(
i,
base_seed = 1253281,
params = list(s = 15, e = 10, w = 40, z = 50, lambda1 = 5, lambda2 = 1.8),
# tweak network
k_min = 2,
k_max = 20,
alpha = 2.4,
rho = 0.4,
r = -0.1,
# retrieve network
return_network = FALSE
) {
# derived seed for this run
seed_i <- base_seed + i
set.seed(seed_i)
# --- network creation ---
degseq <- fdegseq(
n = 100,
alpha = alpha,
k_min = k_min,
k_max = k_max,
dist = "log-normal", #use log-normal
seed = seed_i
)
network <- sample_degseq(degseq, method = "vl")
V(network)$role <- sample(
c(rep("trendsetter", 10), rep("conformist", 90))
)
rewired_network <- frewire_r(network, r, verbose = FALSE, max_iter = 1e5)
final_network <- fswap_rho(rewired_network, rho, verbose = FALSE, max_iter = 1e4)
# --- stats ---
stats <- list(
run = i,
seed = seed_i,
num_nodes = vcount(final_network),
num_edges = ecount(final_network),
avg_degree = mean(degree(final_network)),
sd_degree = sd(degree(final_network)),
net_density = edge_density(final_network),
net_diameter = diameter(final_network, directed = FALSE, unconnected = TRUE),
avg_path_len = average.path.length(final_network, directed = FALSE),
clust_coeff = transitivity(final_network, type = "global"),
assort_deg = assortativity_degree(final_network),
deg_trait_cor = fdegtraitcor(final_network)$cor,
components = components(final_network)$no
)
fplot_graph(final_network, layout = layout_with_fr(final_network))
# --- initial actions ---
V(final_network)$action <- ifelse(V(final_network)$role == "trendsetter", 1, 0)
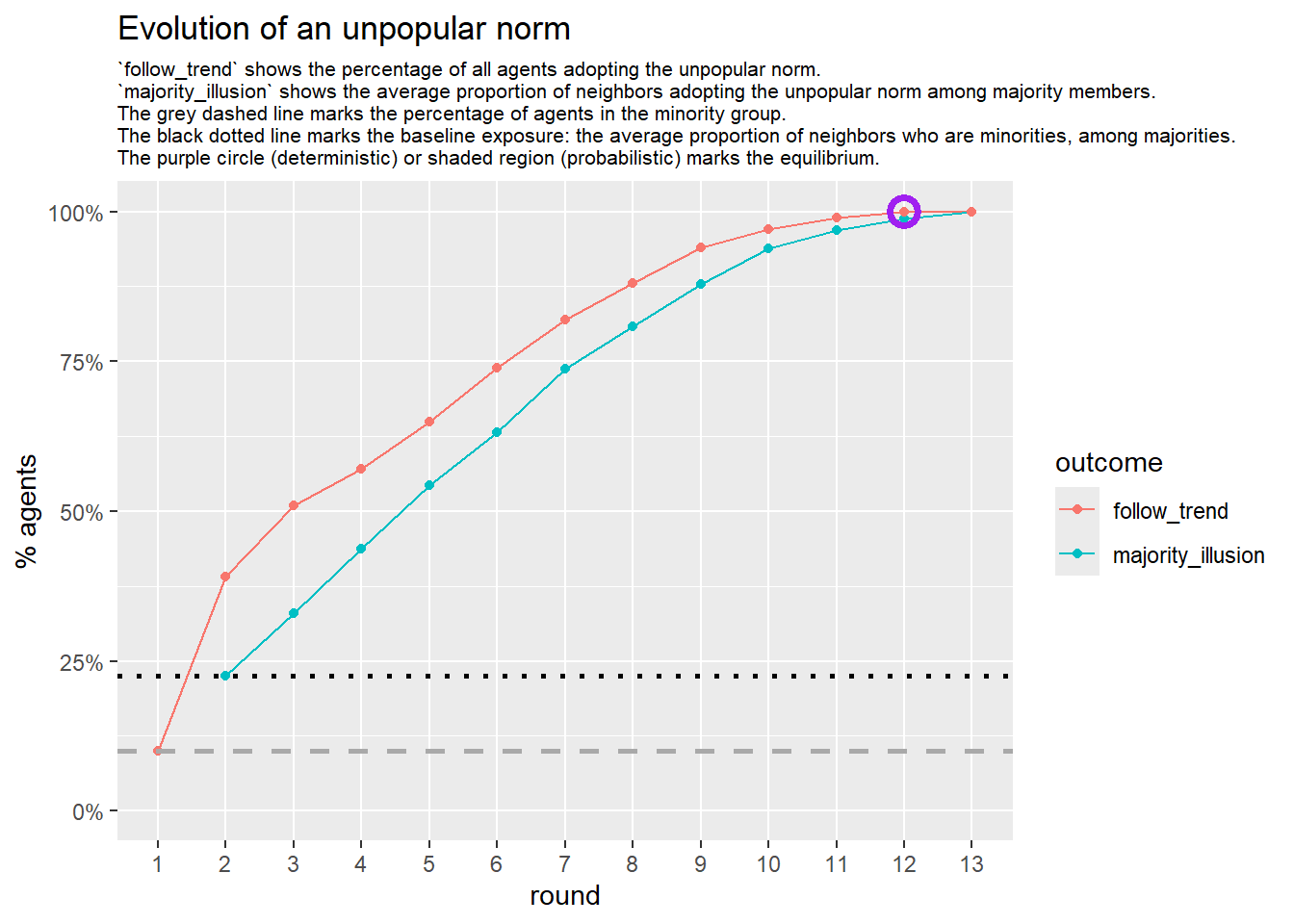
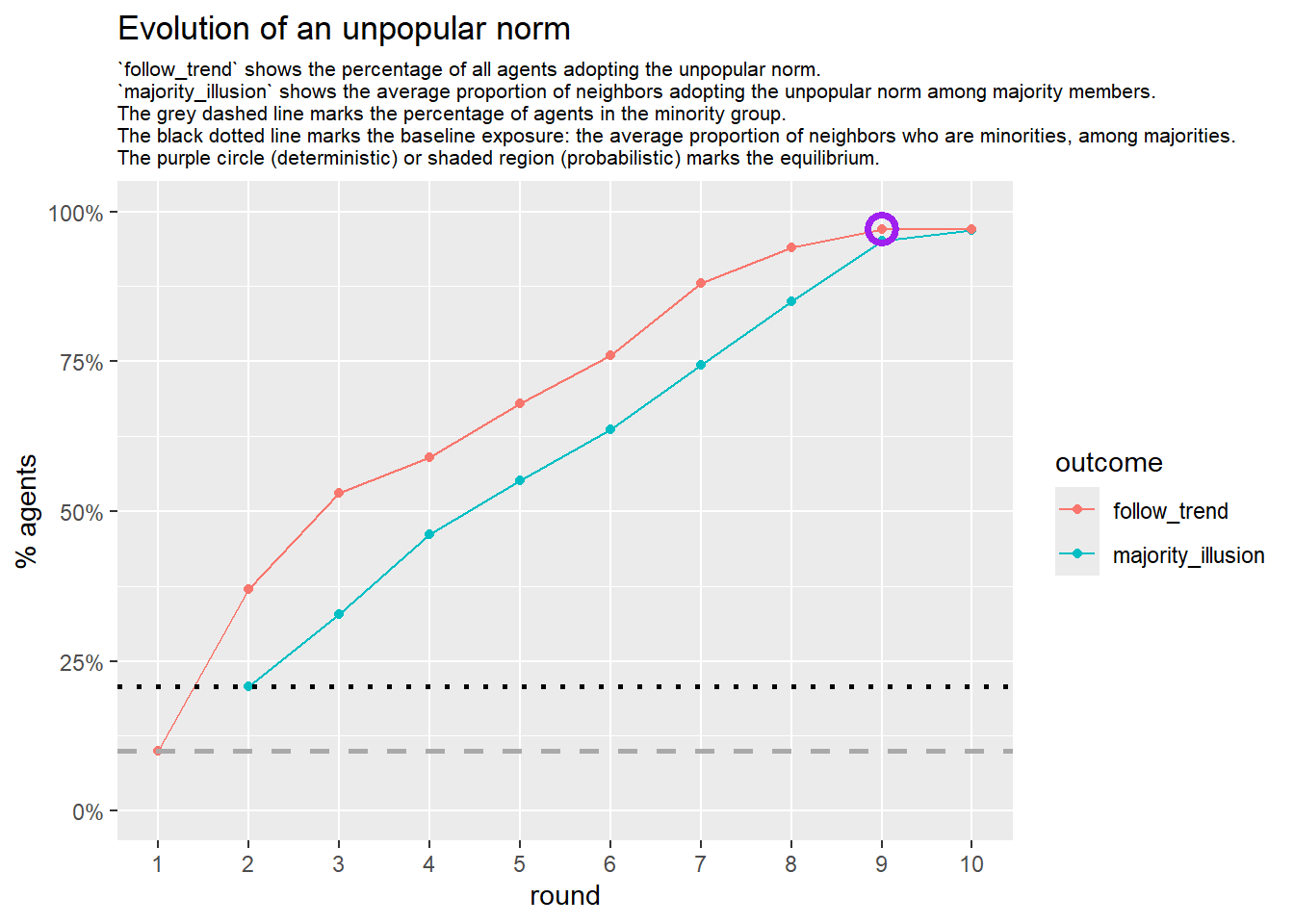
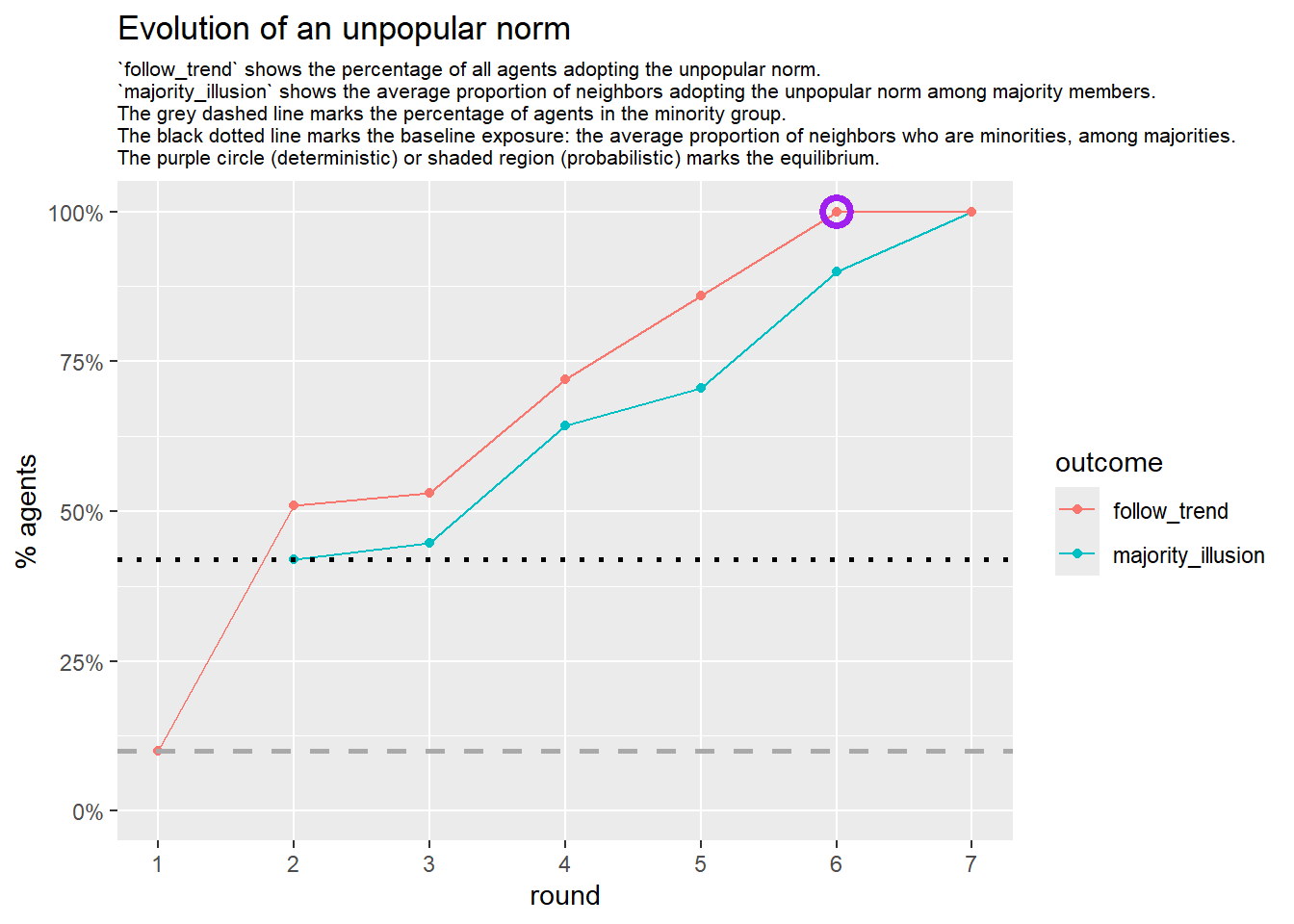
# --- deterministic simulation ---
sim_det <- fabm(
network = final_network,
params = params,
max_rounds = 50,
mi_threshold = 0.49,
choice_rule = "deterministic",
plot = TRUE,
histories = TRUE
)
# generate the gif for the current network
gif_filename <- paste0("./figures/animation_network_", seed_i, ".gif")
gif_path <- fnetworkgif(final_network, sim_det$decision_history, rounds = sim_det$equilibrium$round, output_dir = "./figures")
# rename the gif to match the naming pattern
file.rename(gif_path, gif_filename)
if (!is.null(sim_det$plot)) {
print(sim_det$plot)
}
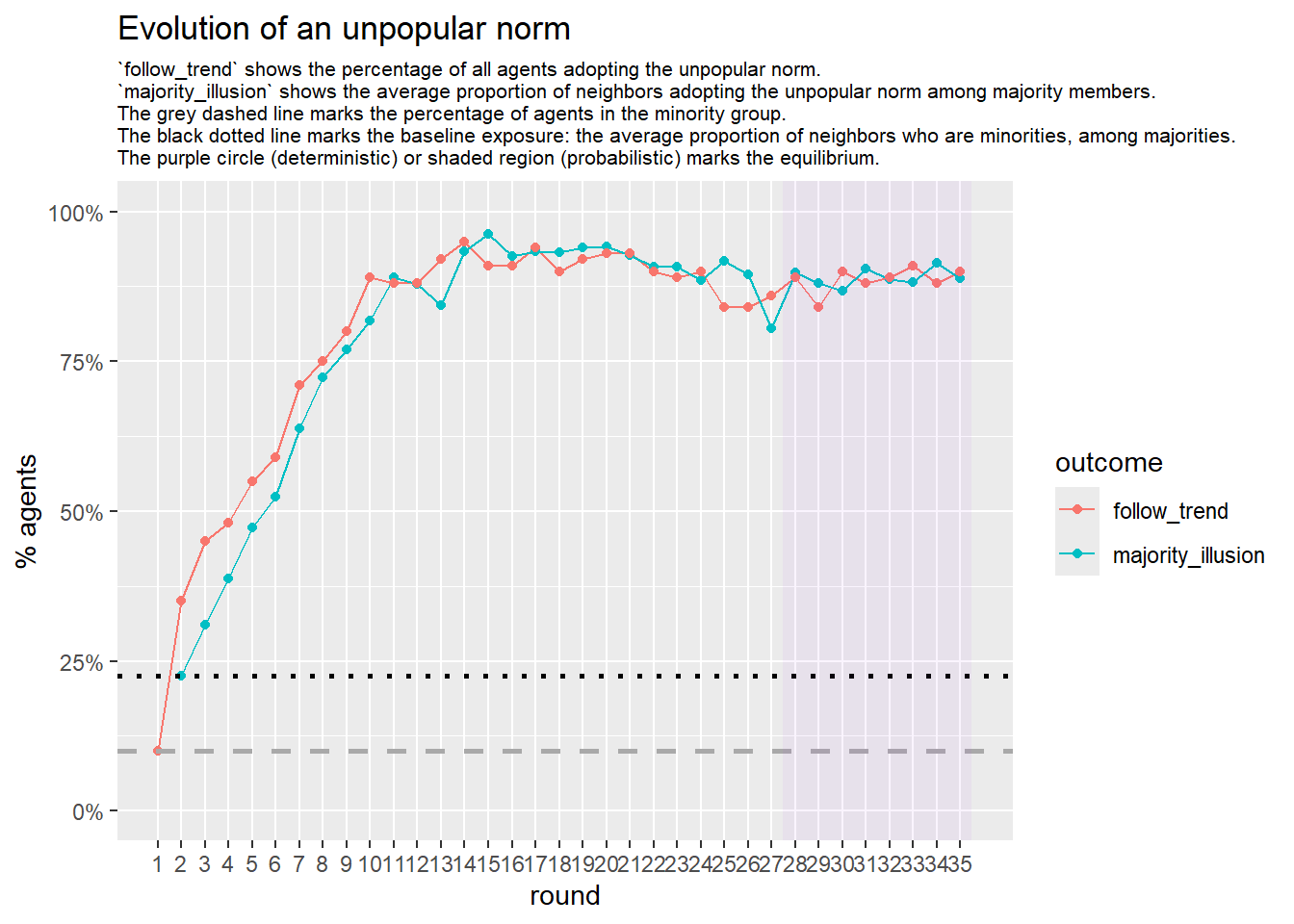
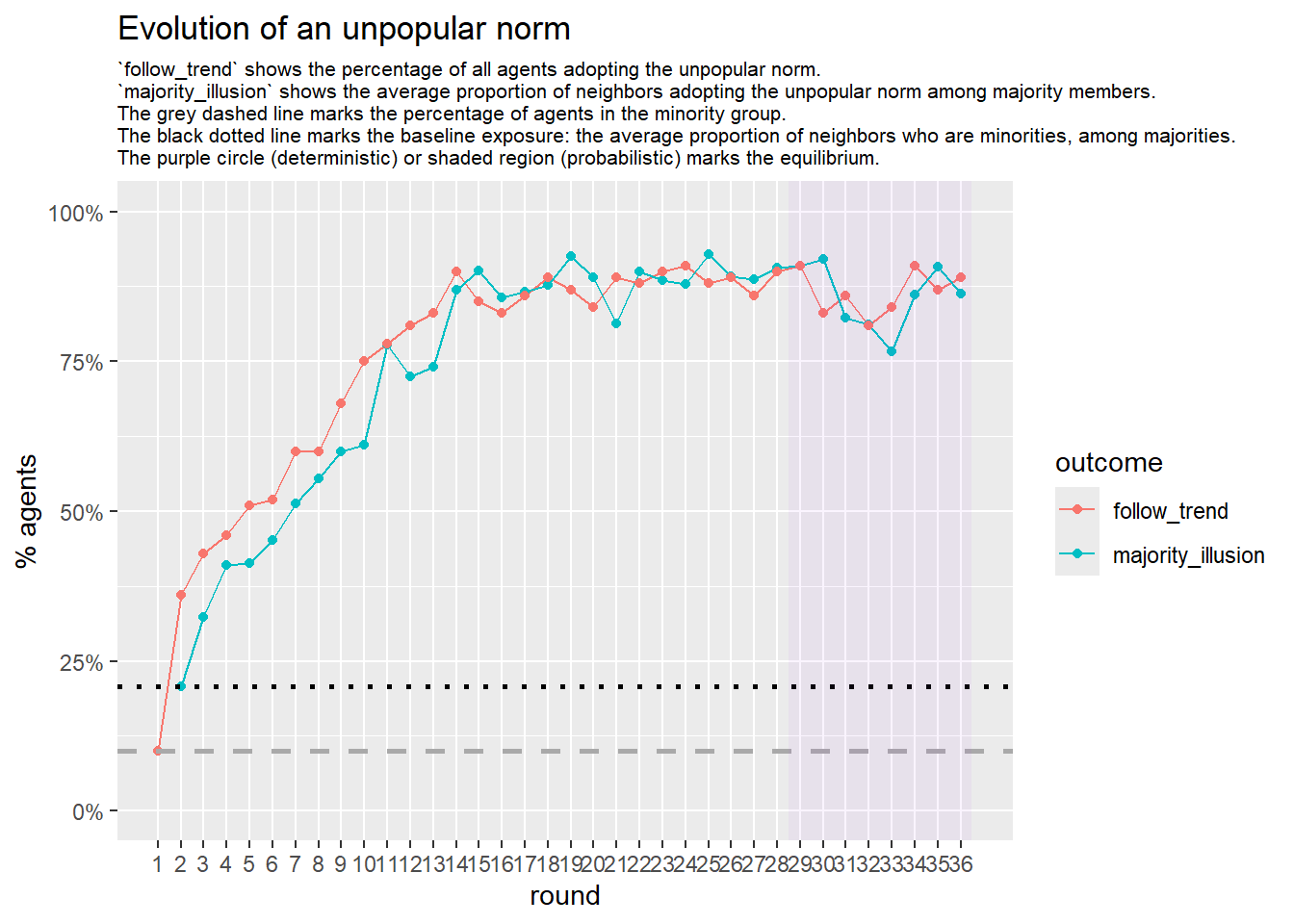
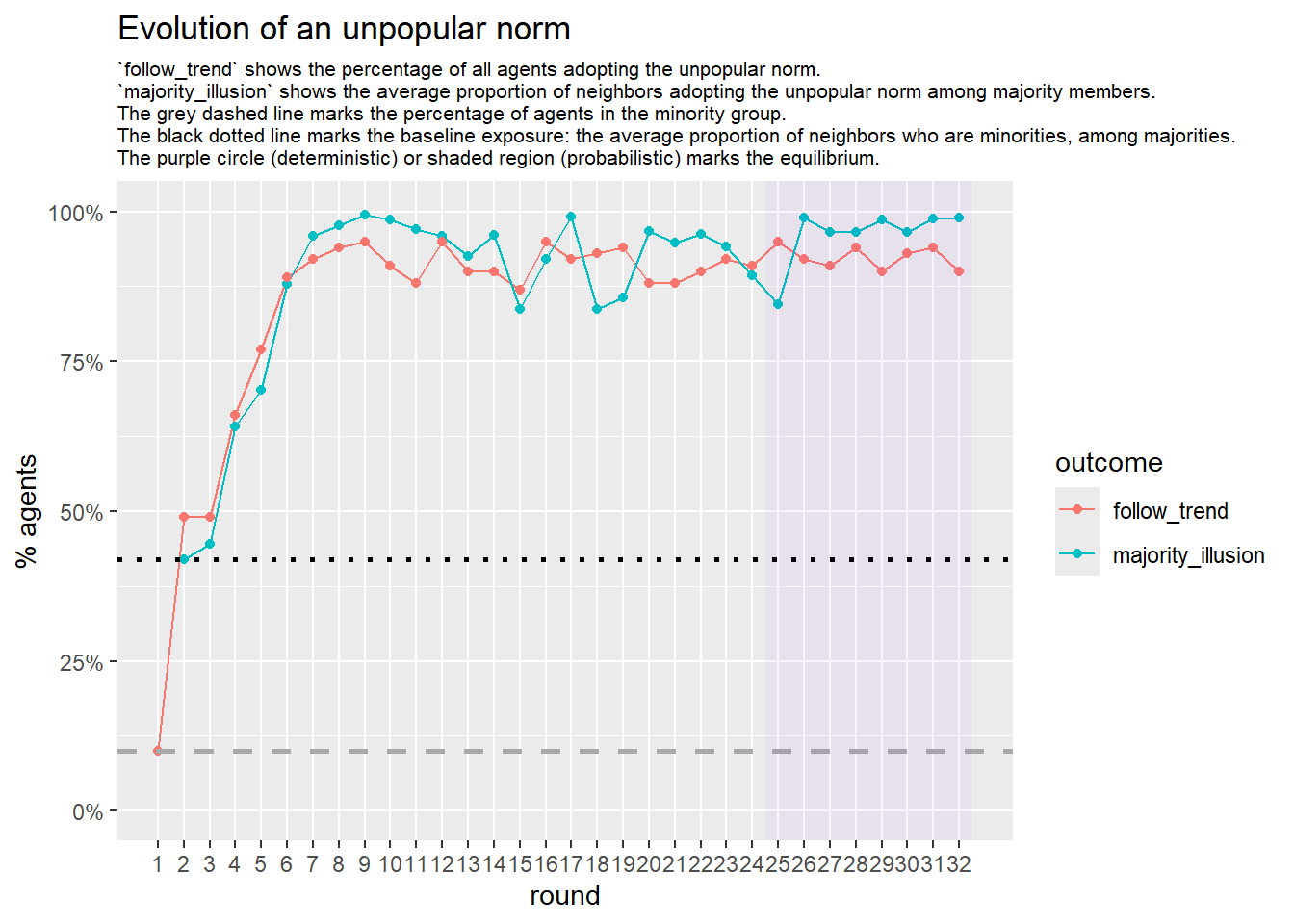
# --- probabilistic simulation ---
sim_prob <- fabm(
network = final_network,
params = params,
max_rounds = 100,
mi_threshold = 0.49,
choice_rule = "probabilistic",
stable_window = 8, # the length of the window of adoption values
required_stable_rounds = 20, # number of windows needed to declare equilibrium
plot = TRUE
)
if (!is.null(sim_prob$plot)) {
print(sim_prob$plot)
}
result <- list(
segregation_det = sim_det$equilibrium$segregation,
segregation_prob = sim_prob$equilibrium$segregation,
stats = stats
)
if (return_network) {
result$network <- final_network
}
result
}test <- run_one_seed(1, k_min = 2, k_max = 20, alpha = 2.4, rho = 0.4, r = -0.1, return_network = TRUE)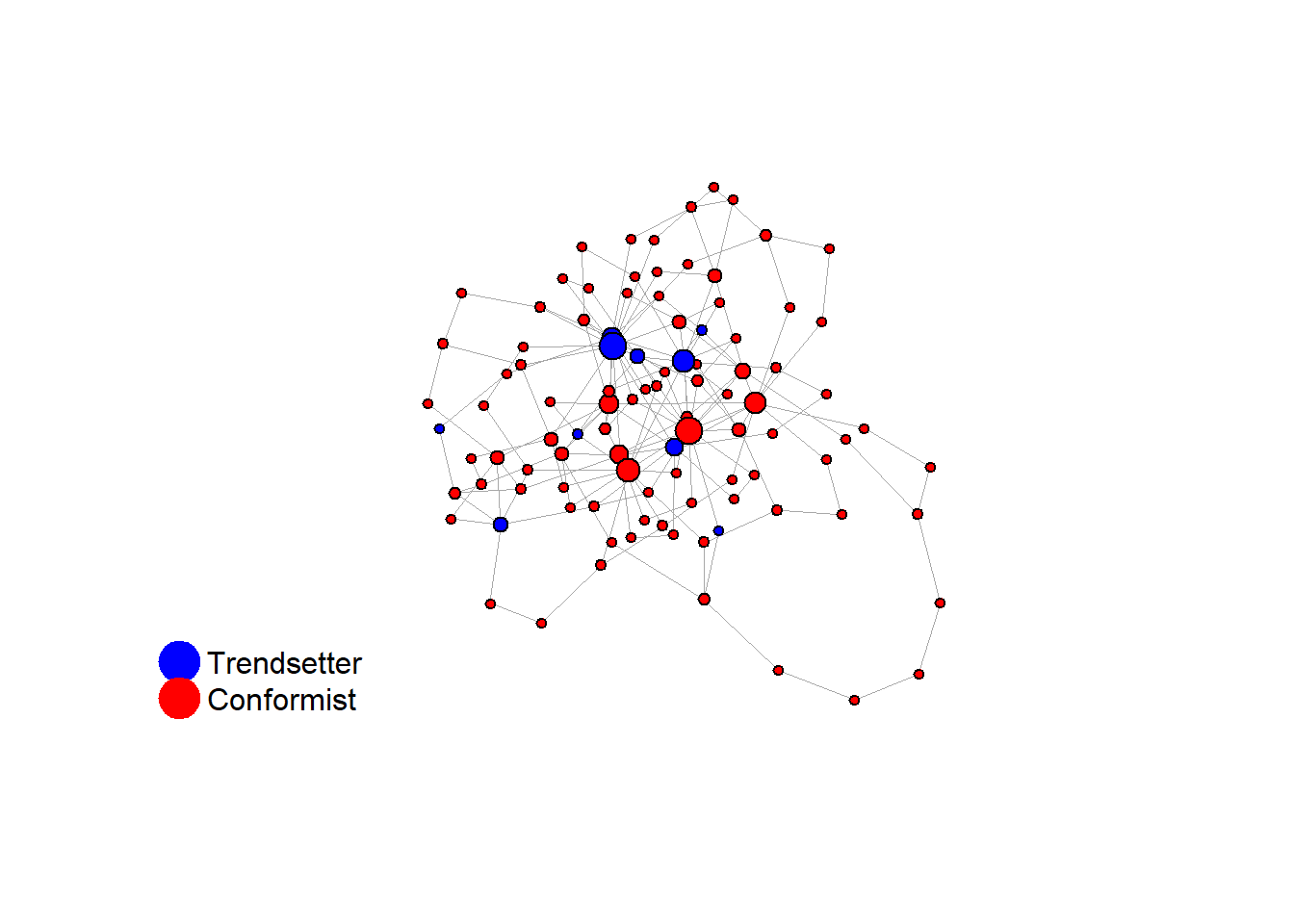
base = 1253281
seed = base + 1knitr::include_graphics(paste0("./figures/animation_network_", seed, ".gif"))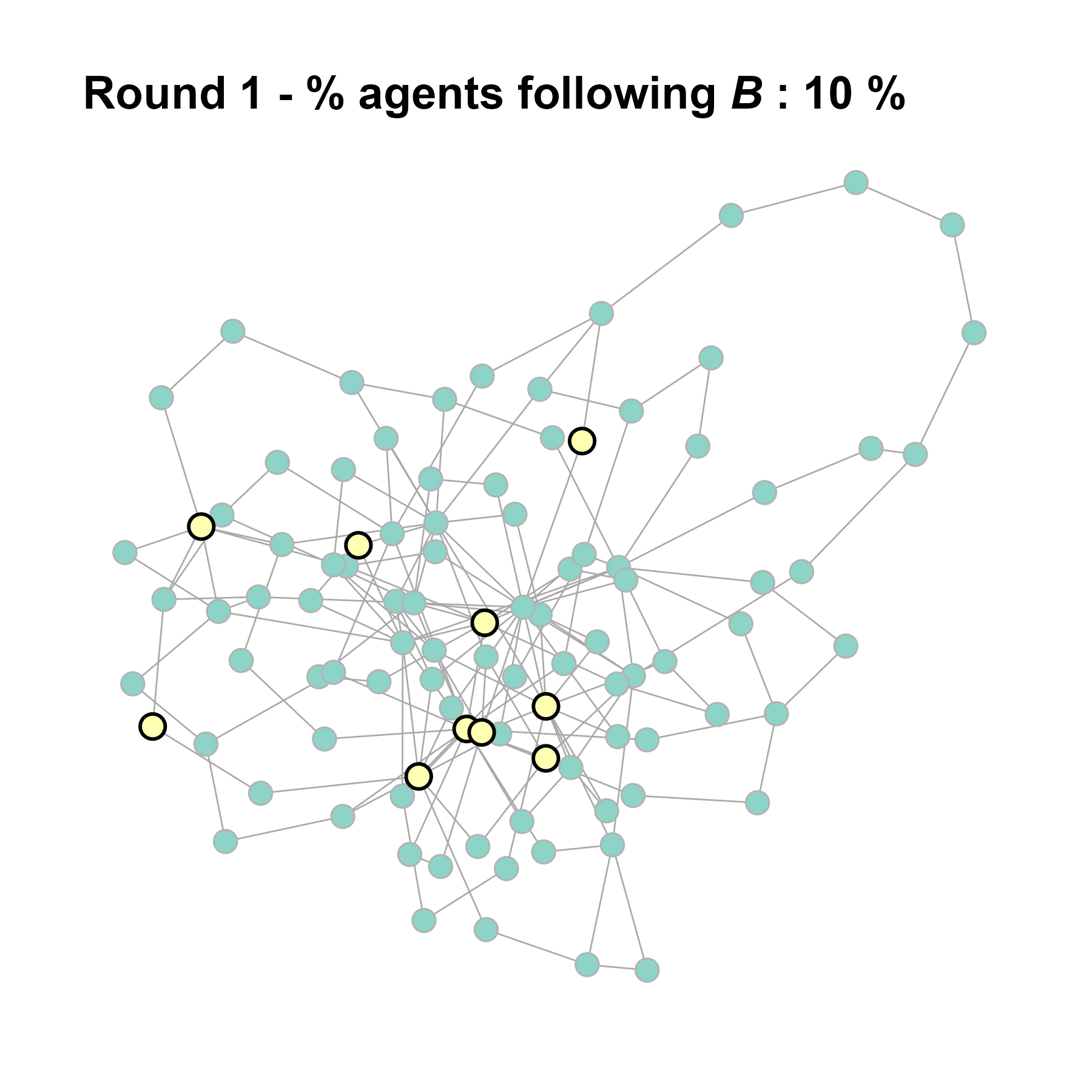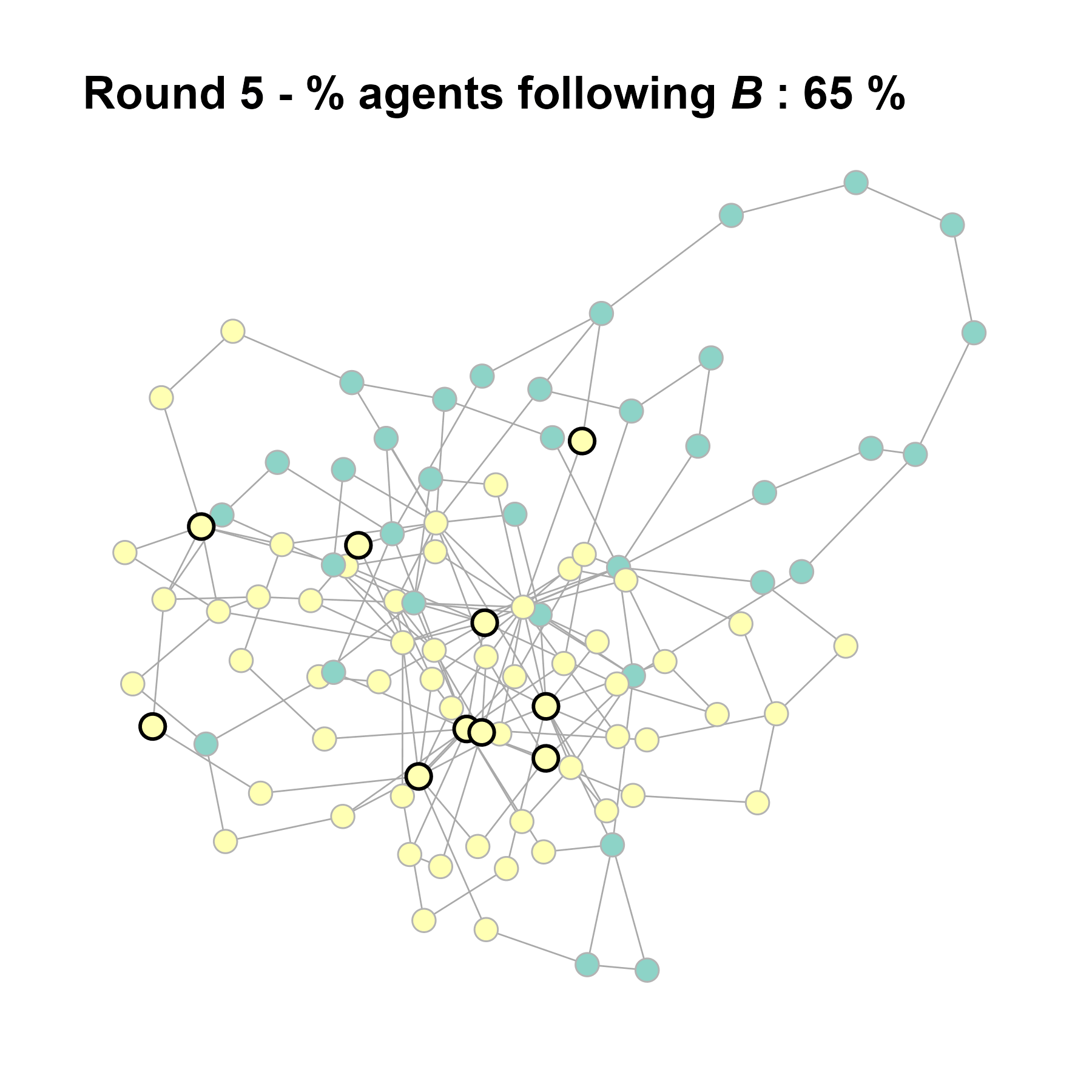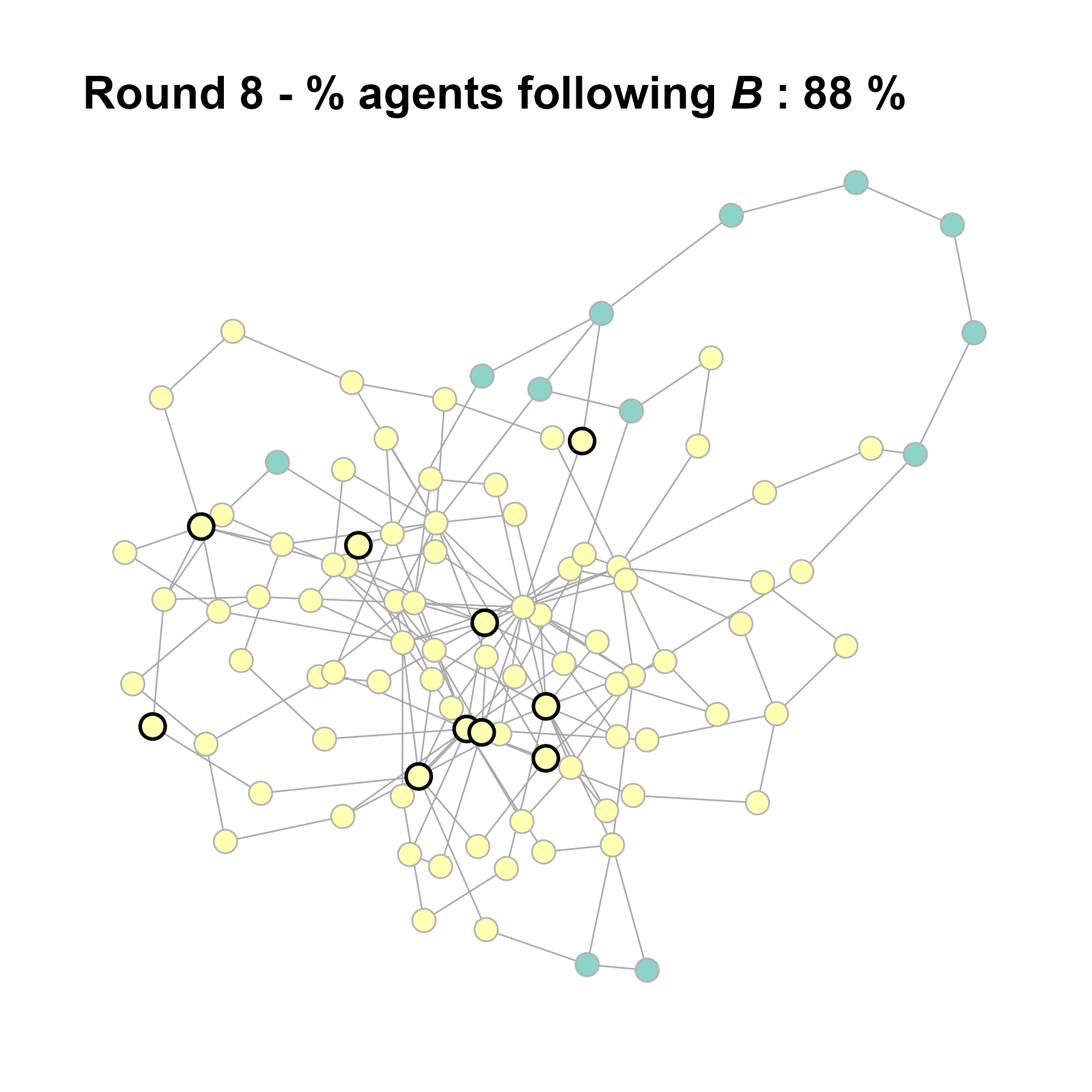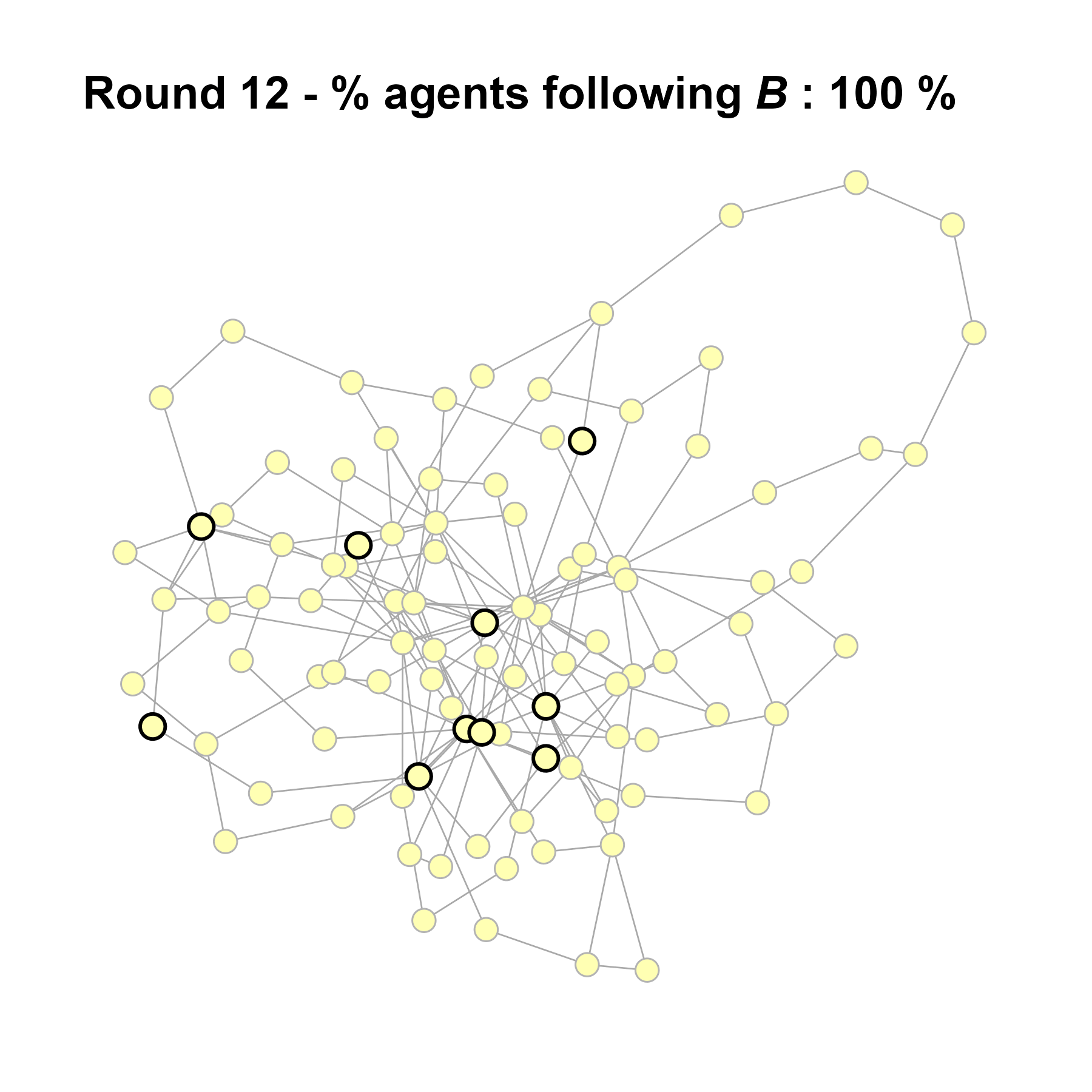
test <- run_one_seed(23, k_min = 2, k_max = 80, alpha = 2.4, rho = 0.4, r = -0.1, return_network = TRUE)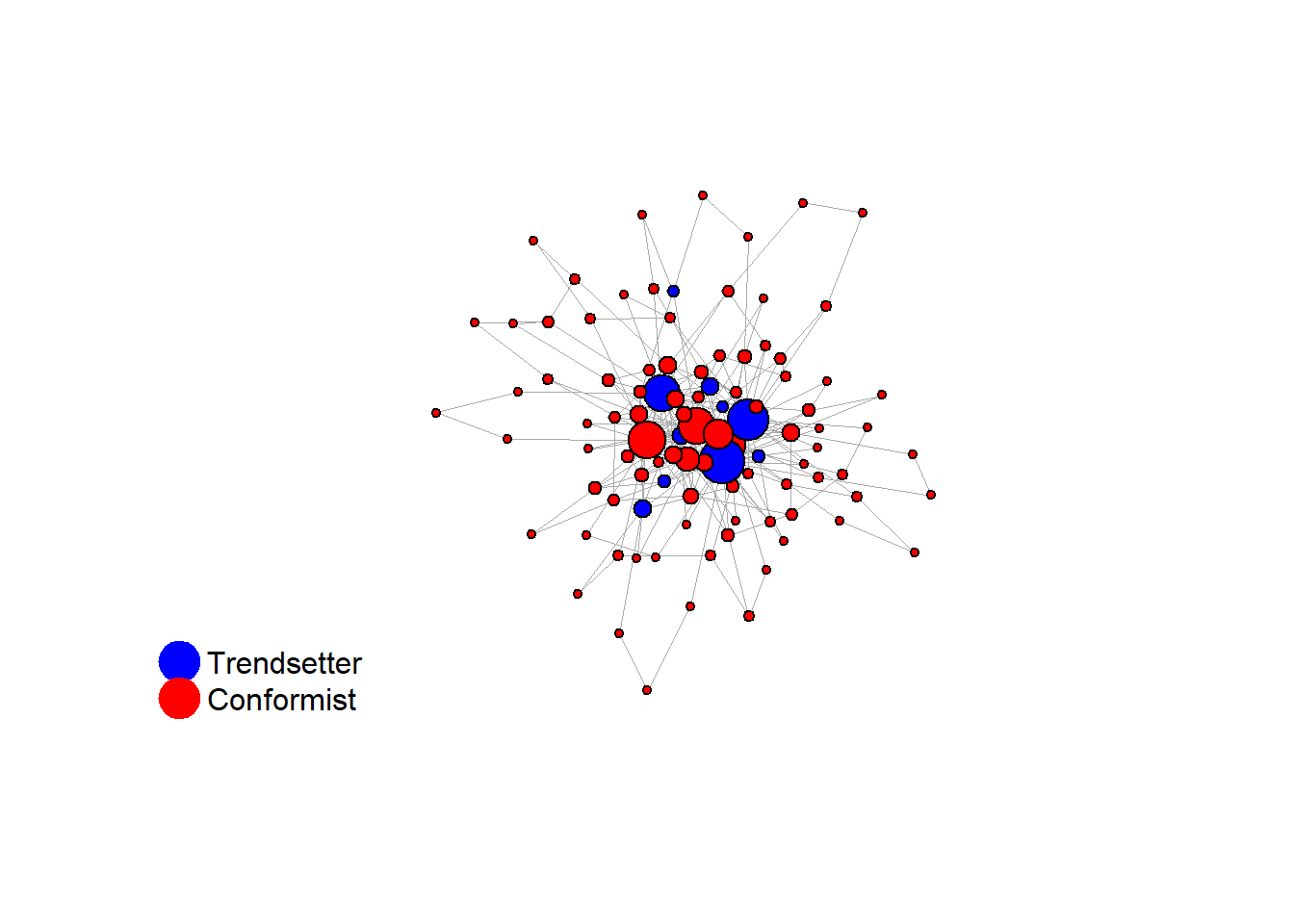
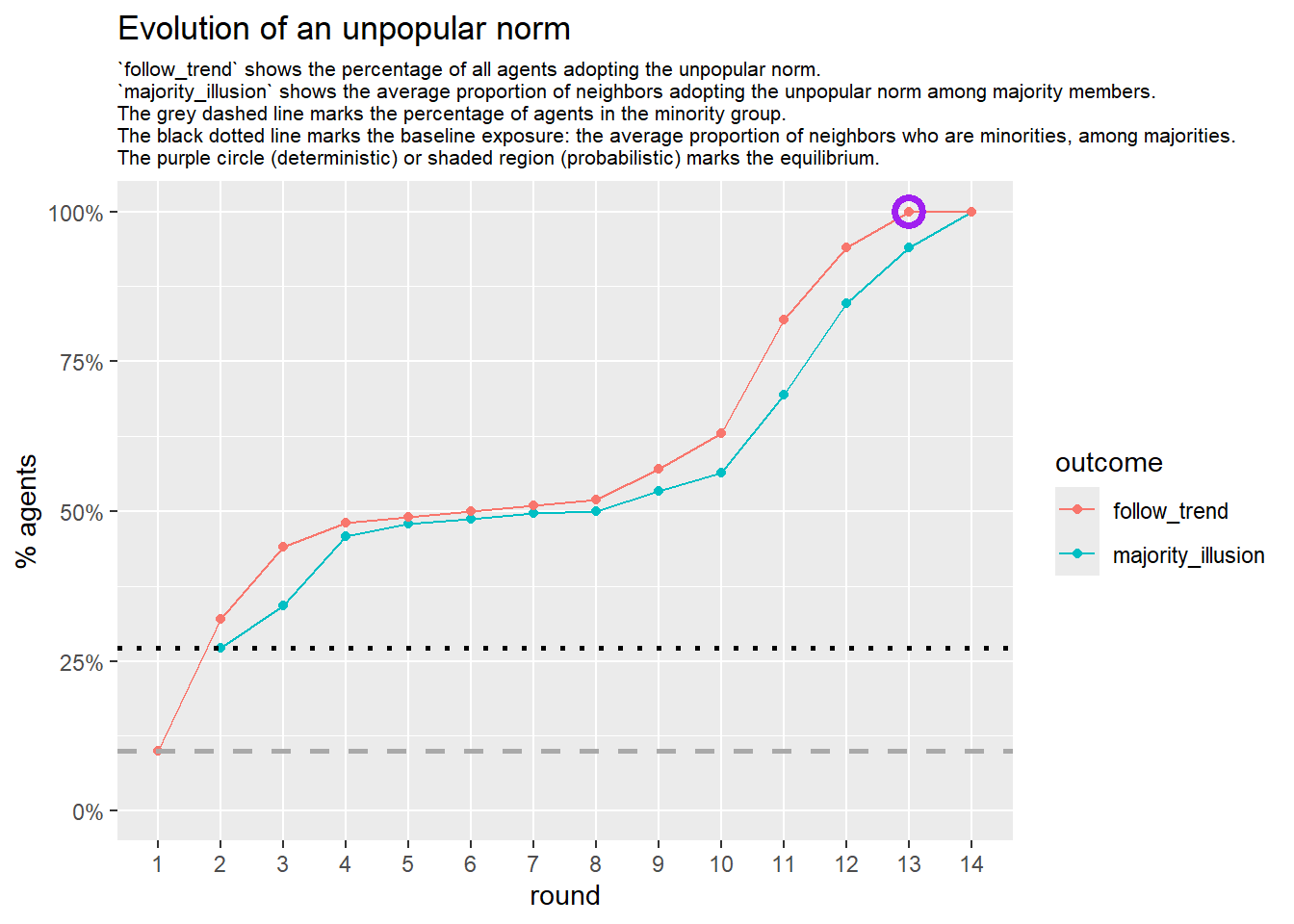
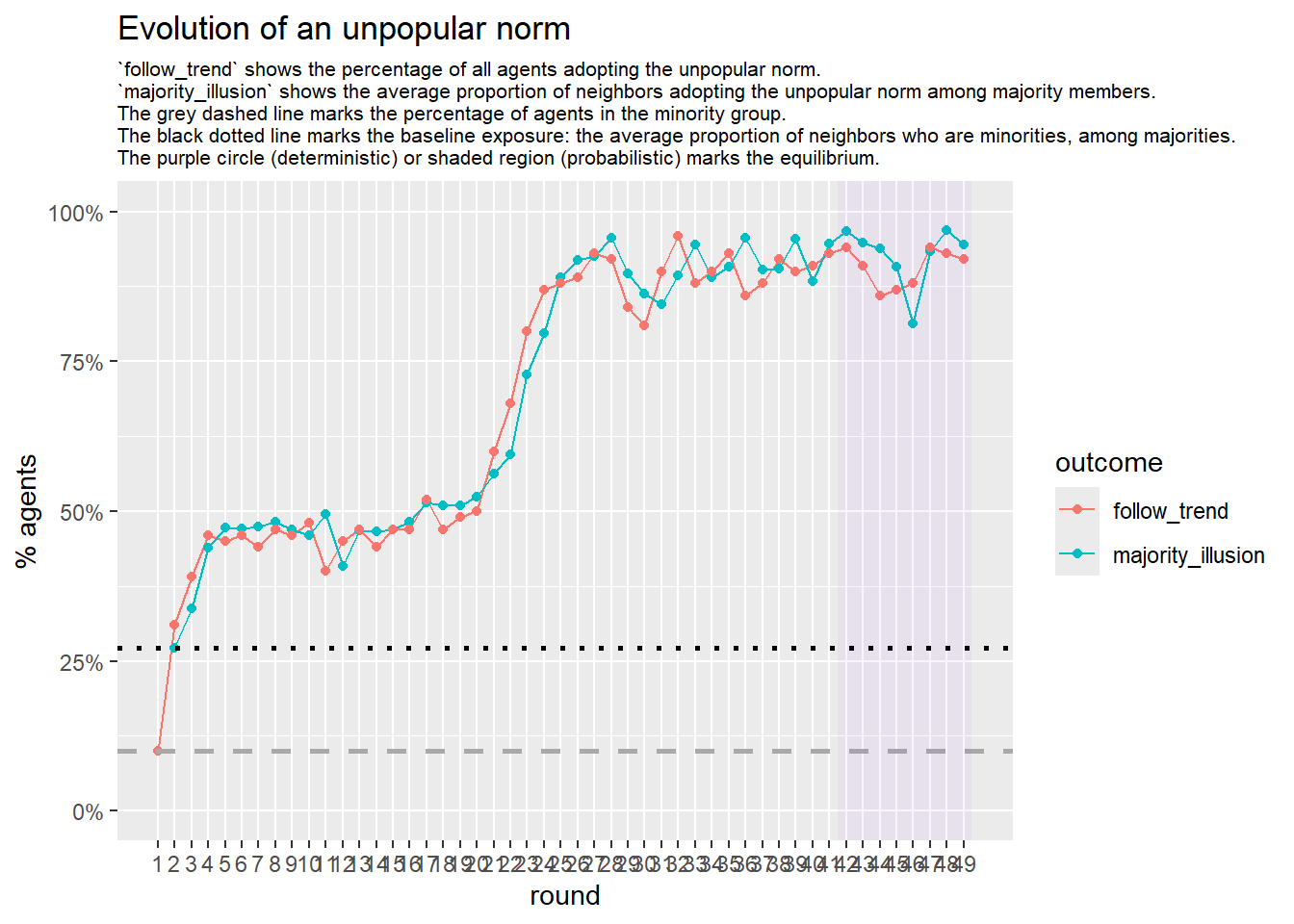
seed = seed + 22
# 22
# 23, 80 #typical s-curve (traction among susceptibles... spraeding in the middle; targeting
# susceptibles not direclty exposed.)
# let's check the distribution of distances to 'trendsetters', among 'conformists'
g <- test$network
trend <- which(V(g)$role == "trendsetter")
conf <- which(V(g)$role == "conformist")
# distances: rows = sources (trendsetters), cols = all vertices
D <- distances(g, v = trend, to = V(g), mode = "all")
Dc <- D[, conf] #keep just conformists
# calculate (a) the distance to the nearest trendsetter and (b) the average distance to
# trendsetters, across conformists
dists <- data.frame(shortest = apply(Dc, 2, min), average = apply(Dc, 2, function(x) mean(x[is.finite(x)])))
dists_long <- pivot_longer(dists, cols = c(shortest, average), names_to = "type", values_to = "distance_to_seed")
# plot the distribution
ggplot(dists_long, aes(x = distance_to_seed, fill = type)) + geom_histogram(bins = 20, alpha = 0.5, position = "identity") +
theme_minimal()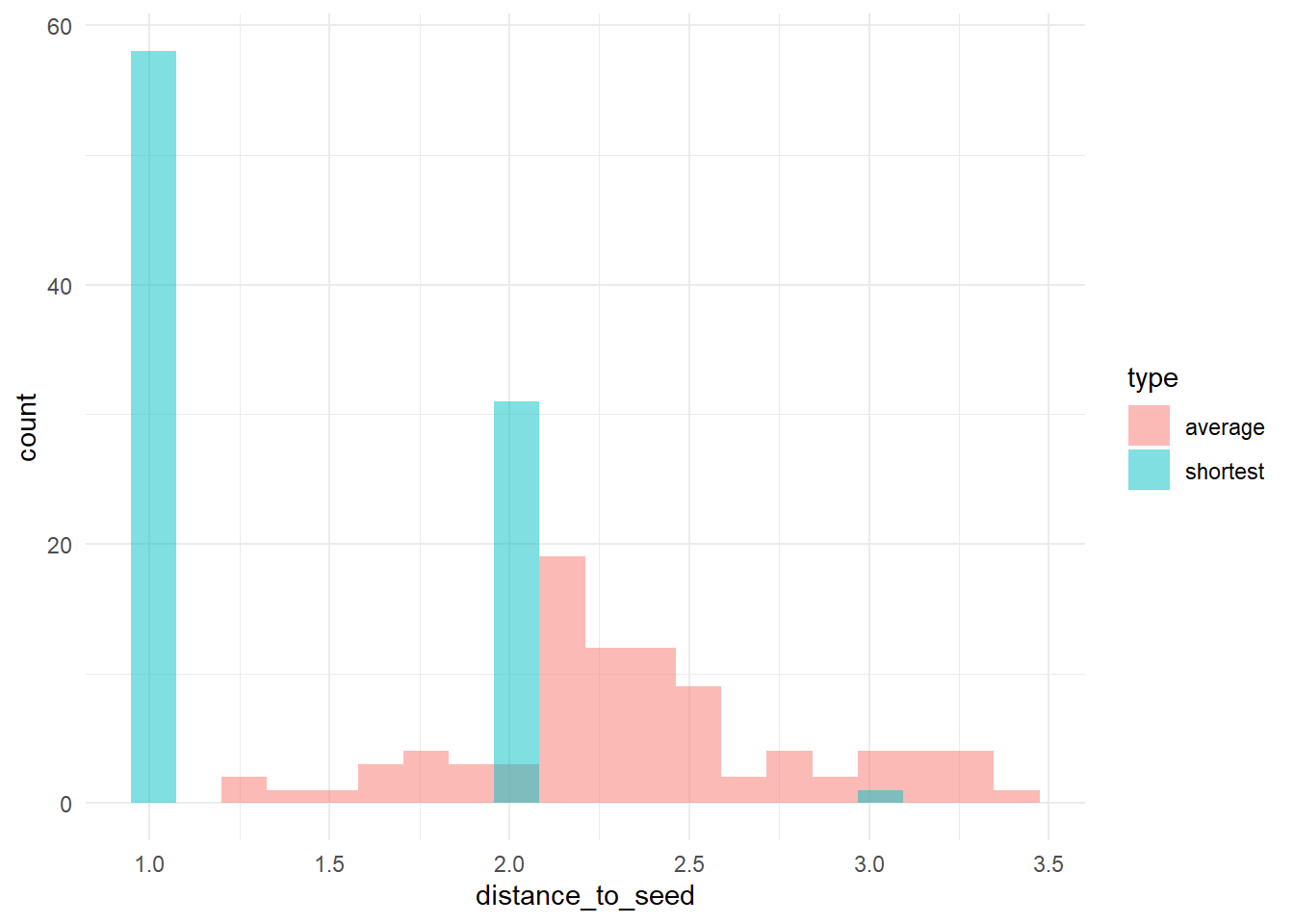
test$stats#> $run
#> [1] 23
#>
#> $seed
#> [1] 1253304
#>
#> $num_nodes
#> [1] 100
#>
#> $num_edges
#> [1] 268
#>
#> $avg_degree
#> [1] 5.36
#>
#> $sd_degree
#> [1] 5.919528
#>
#> $net_density
#> [1] 0.05414141
#>
#> $net_diameter
#> [1] 5
#>
#> $avg_path_len
#> [1] 2.763232
#>
#> $clust_coeff
#> [1] 0.1984154
#>
#> $assort_deg
#> [1] -0.1098723
#>
#> $deg_trait_cor
#> [1] 0.4210629
#>
#> $components
#> [1] 1table(degree(test$network))#>
#> 2 3 4 5 6 7 9 14 19 22 24 25 28 31
#> 37 18 12 9 4 2 10 1 1 1 2 1 1 1# identify trendsetters
ids <- which(V(test$network)$role == "trendsetter")
# check their centrality
sort(as.numeric(cbind(degree(test$network), V(test$network)$role)[ids]))#> [1] 4 4 5 5 9 9 9 24 28 31knitr::include_graphics(paste0("./figures/animation_network_", seed, ".gif"))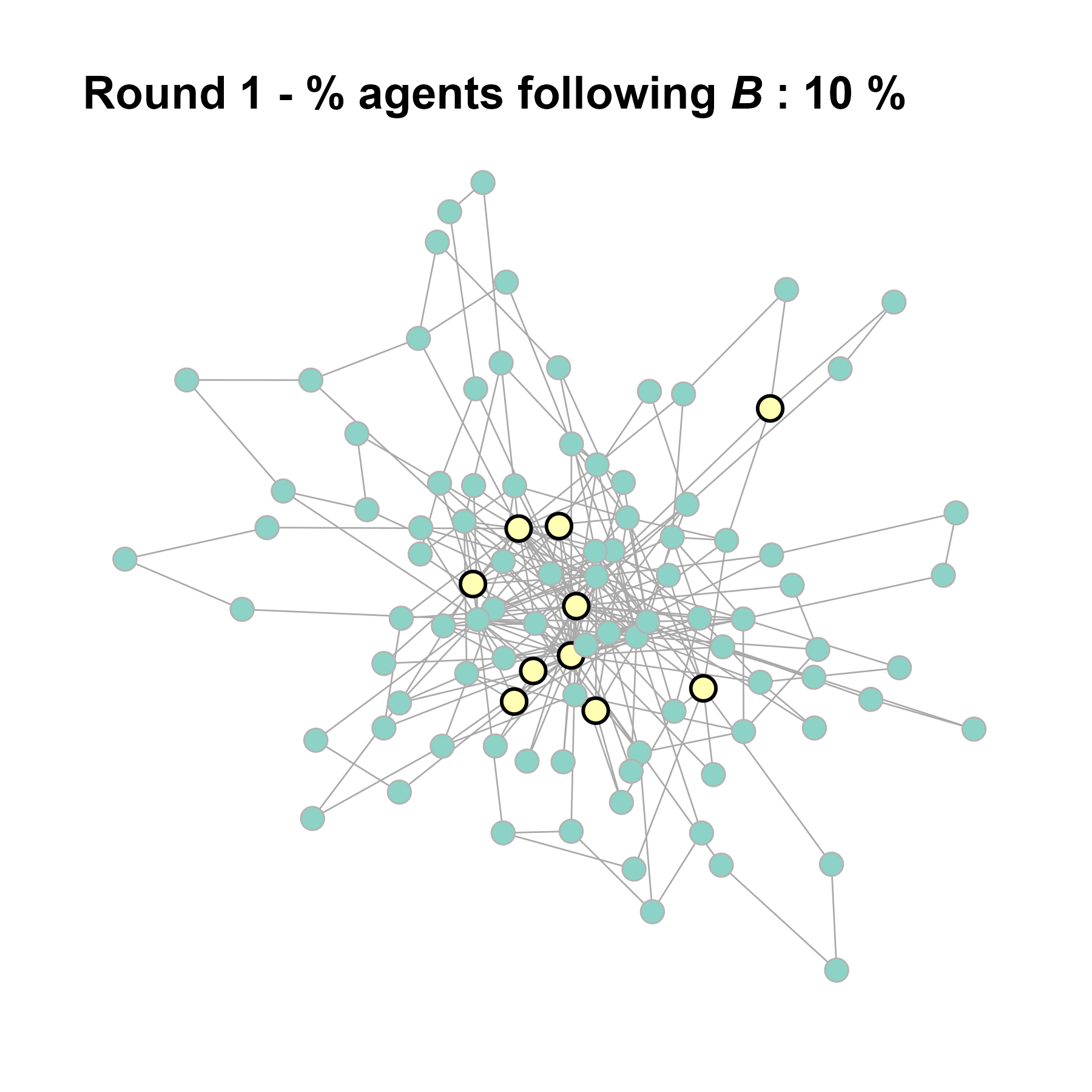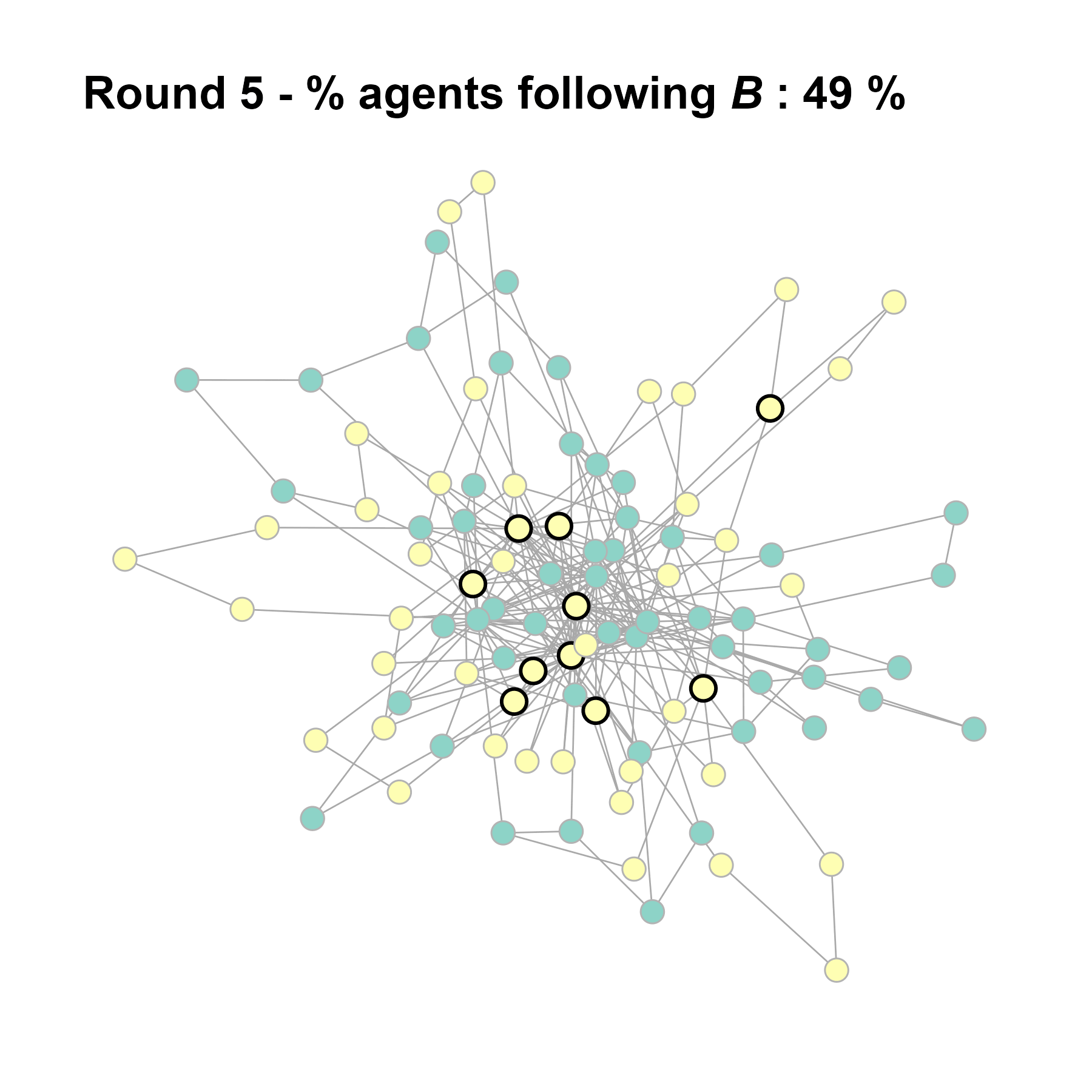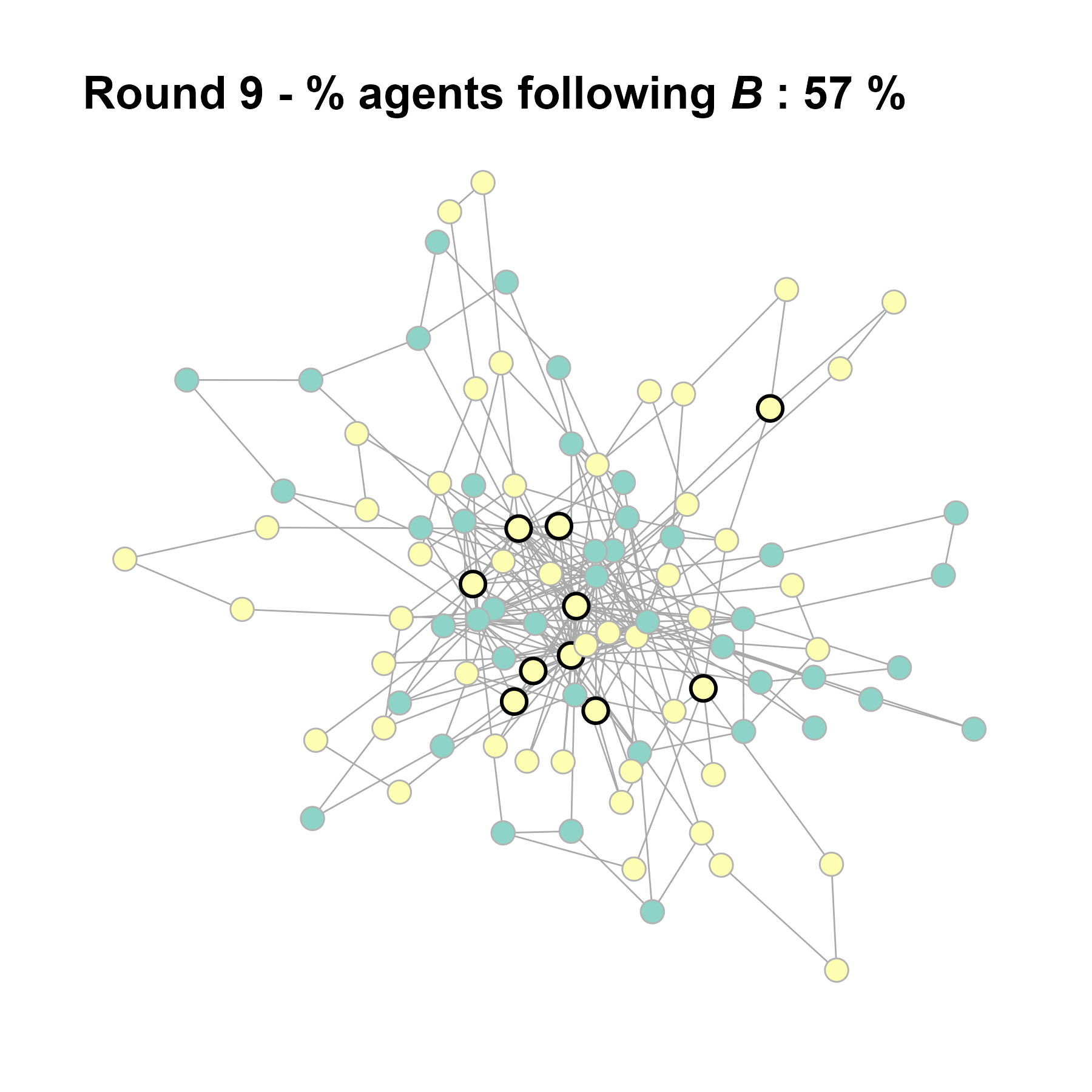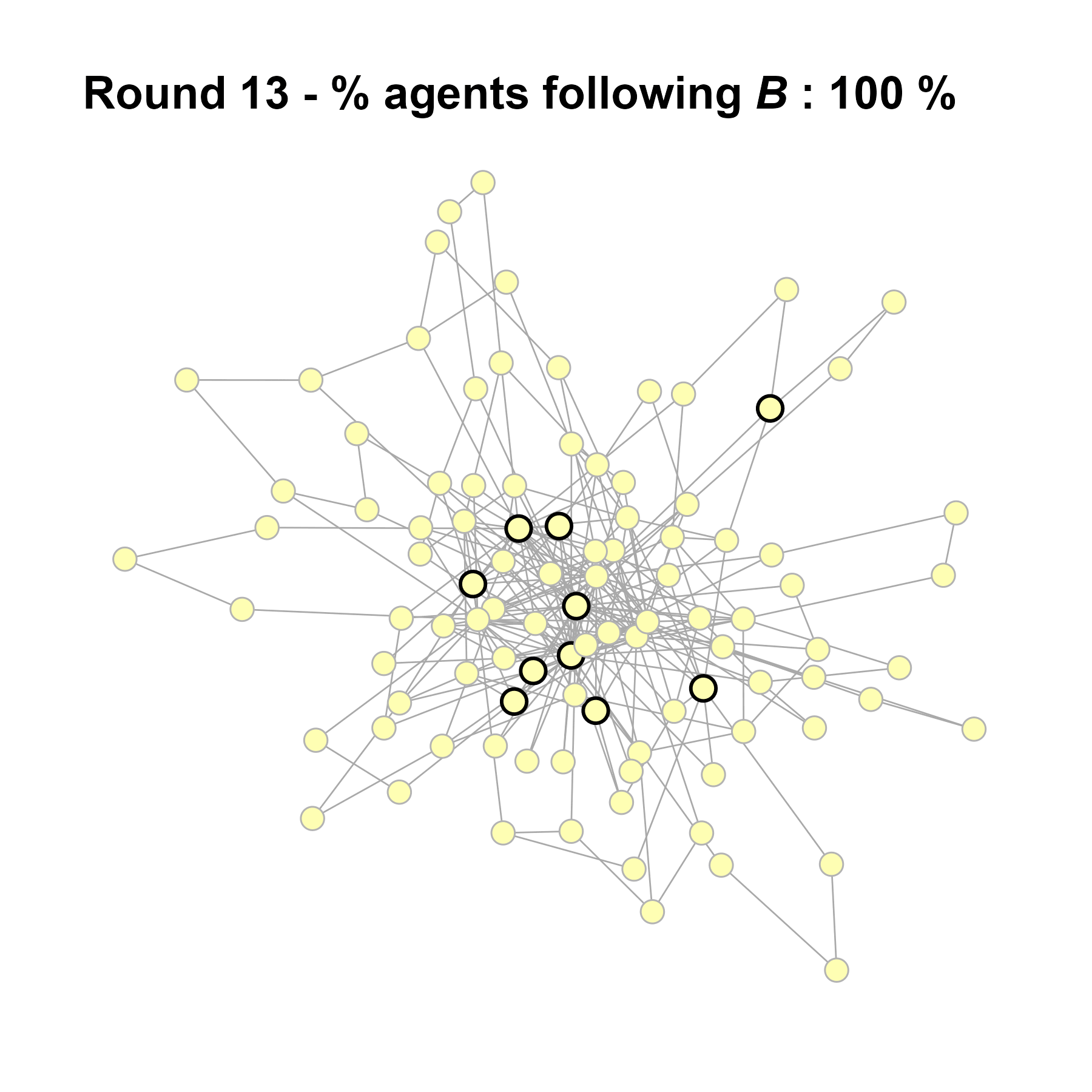
# use this as the network structure for an otree session:
# cbind(degree(test$network),V(test$network)$role)
# convert to adjacency matrix
adj_matrix <- as.matrix(as_adjacency_matrix(test$network))
# get roles
role_vector <- ifelse(V(test$network)$role == "trendsetter", 1, 0)
# create a list to store the network data
net <- list(adj_matrix = adj_matrix, role_vector = role_vector)
# save the list as a JSON file
write_json(net, "network_test_n100.json")test <- run_one_seed(3, k_min = 2, k_max = 20, alpha = 2.4, rho = 0.4, r = -0.1)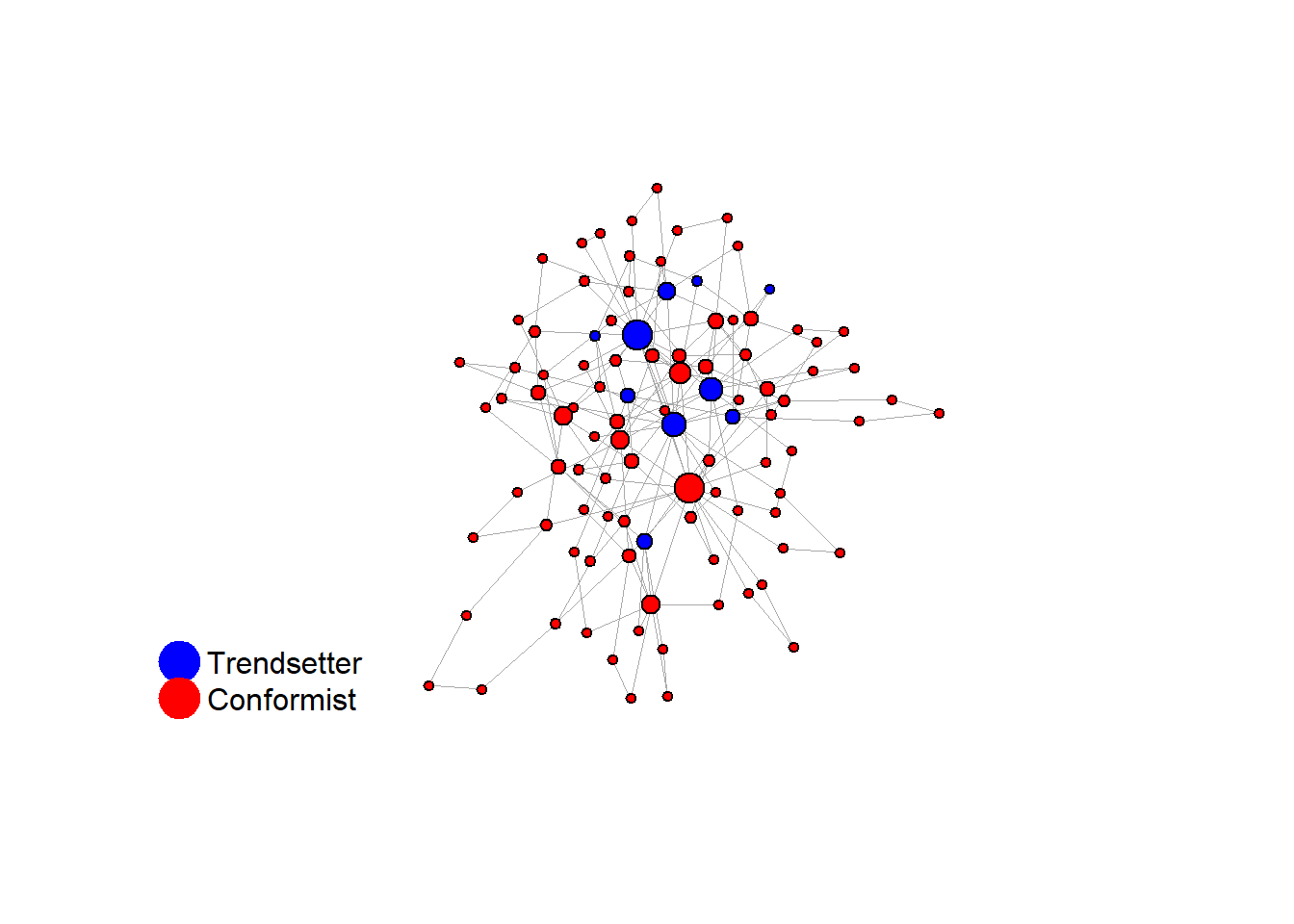
seed = seed - 21
seed = seed + 1knitr::include_graphics(paste0("./figures/animation_network_", seed, ".gif"))
test <- run_one_seed(4, k_min = 2, k_max = 20, alpha = 2.4, rho = 0.4, r = -0.1)
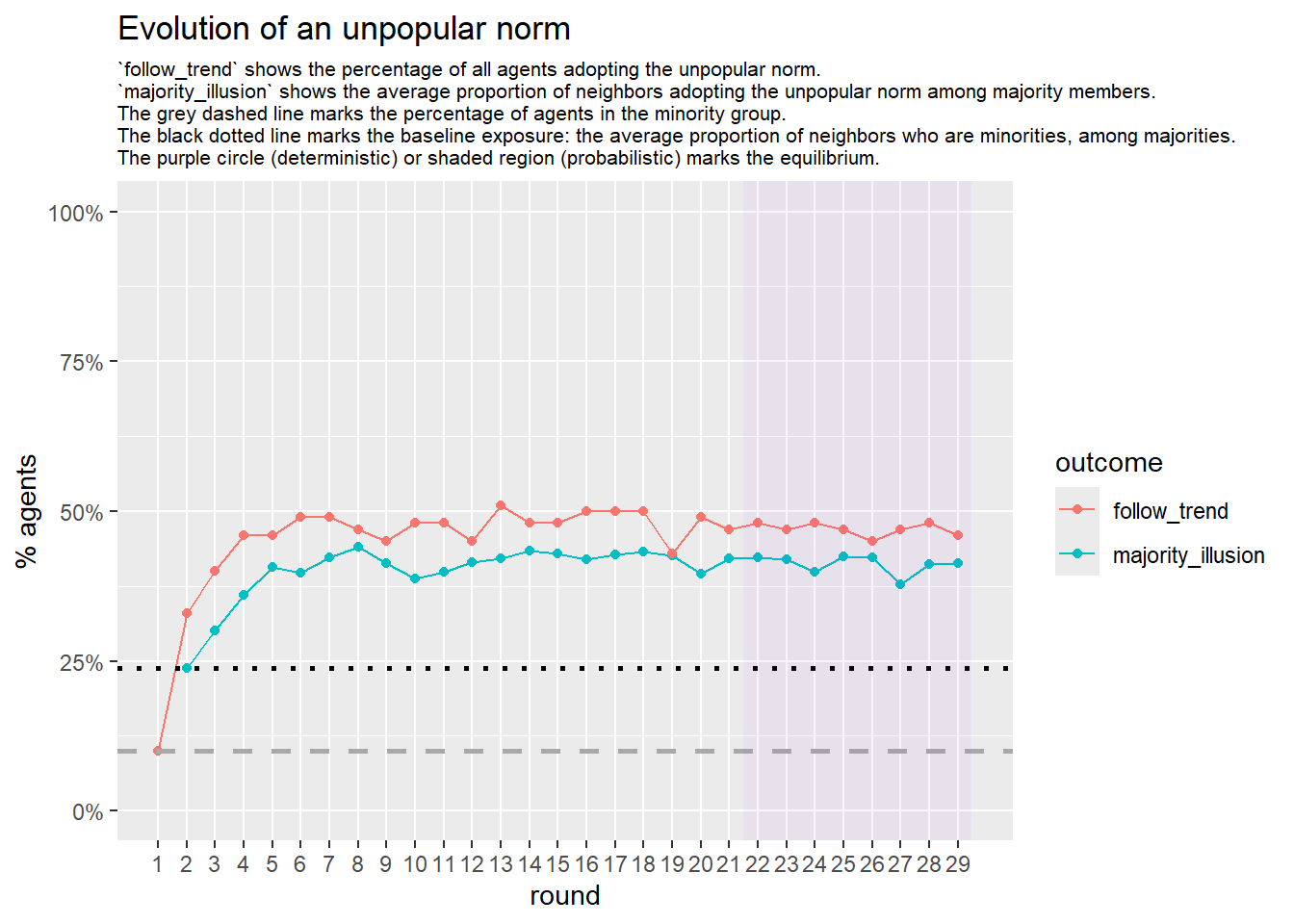
seed = seed + 1knitr::include_graphics(paste0("./figures/animation_network_", seed, ".gif"))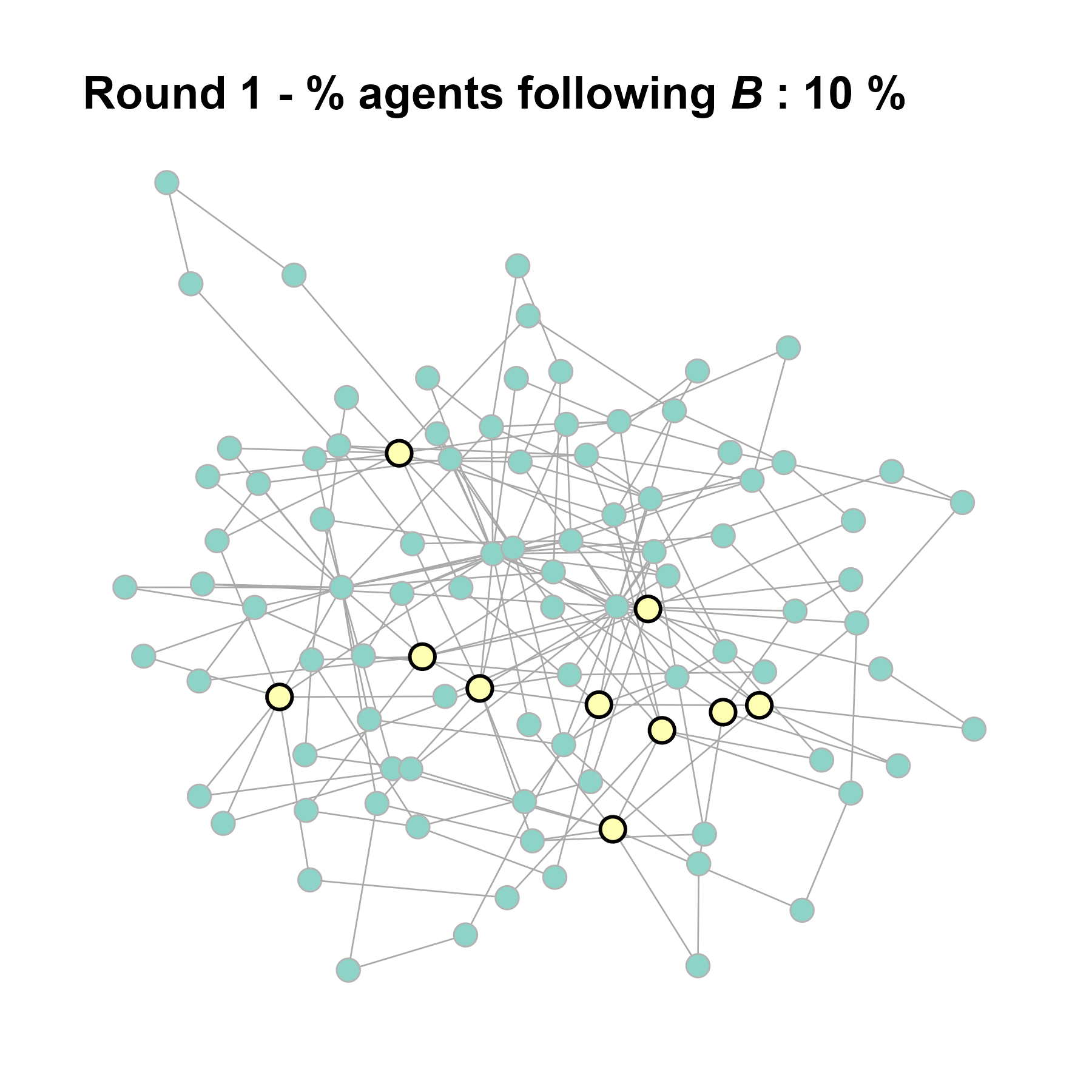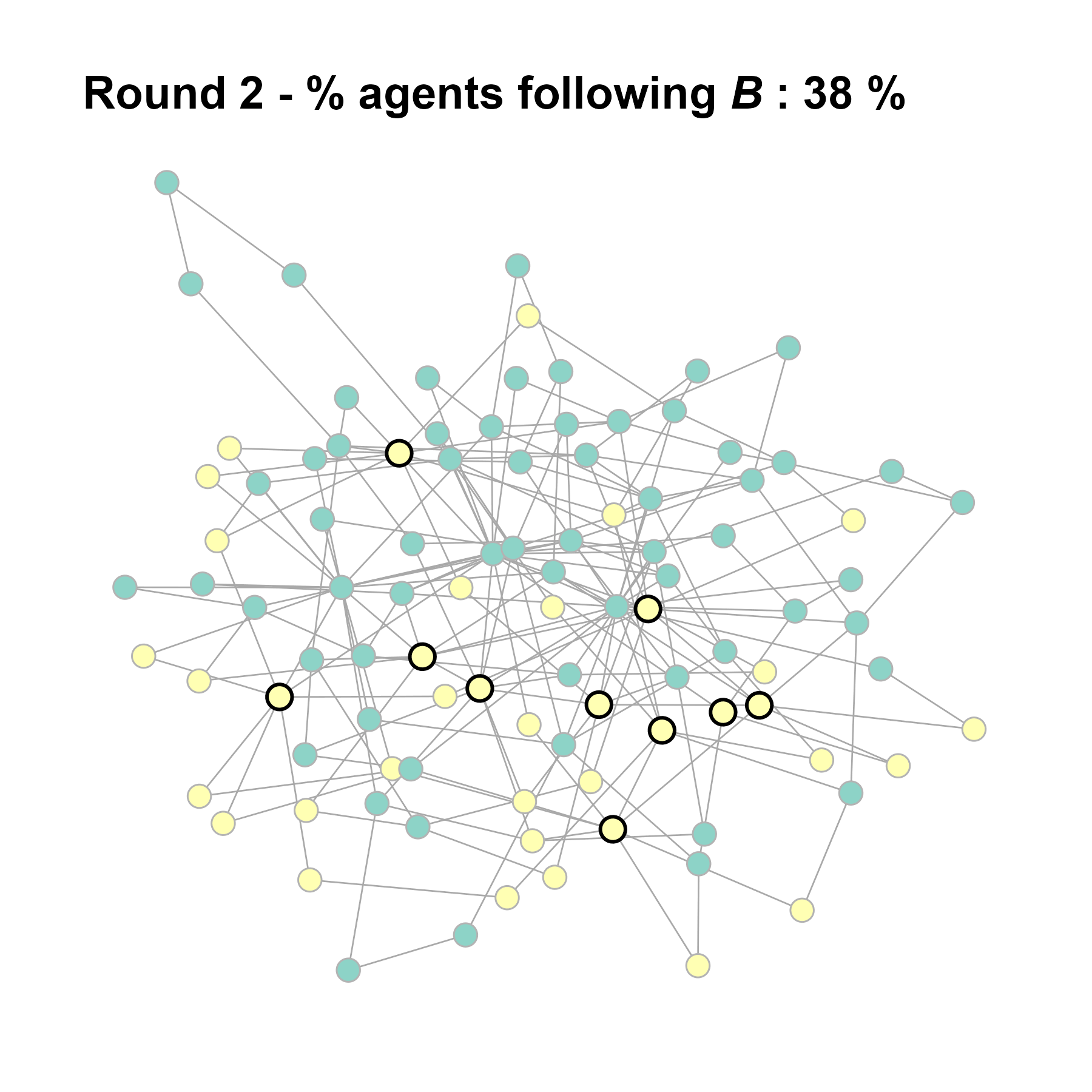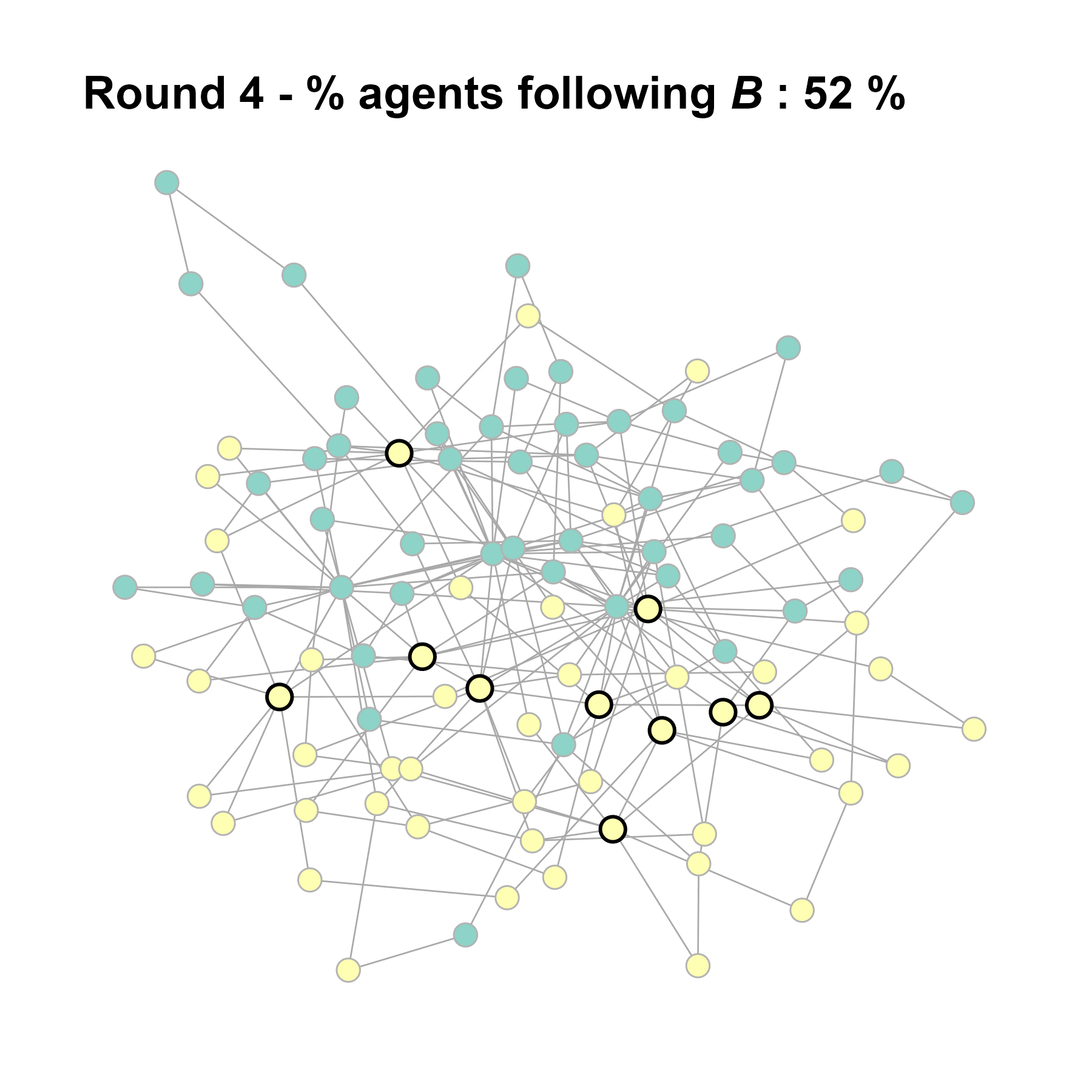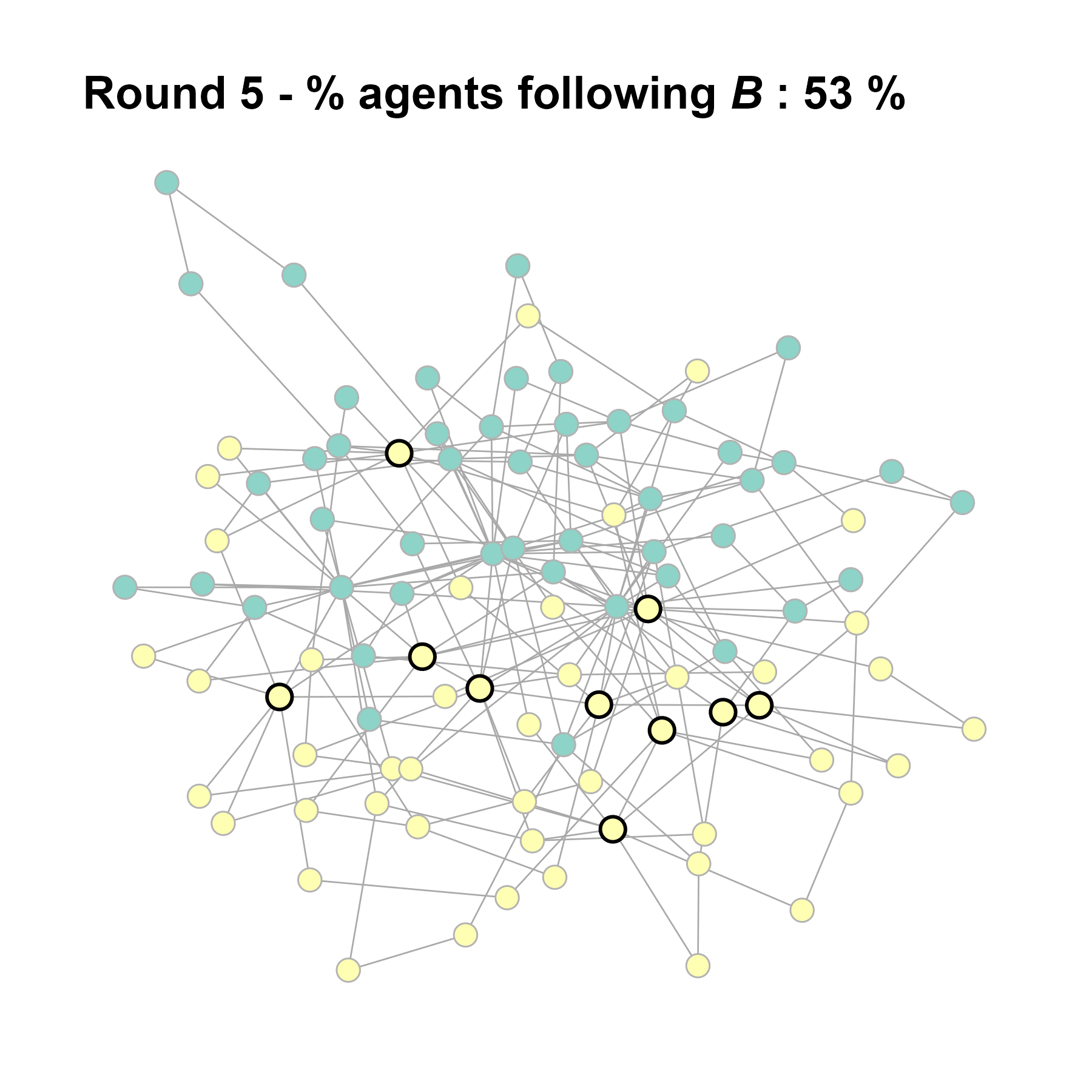
test <- run_one_seed(5, k_min = 2, k_max = 20, alpha = 2.4, rho = 0.4, r = -0.1)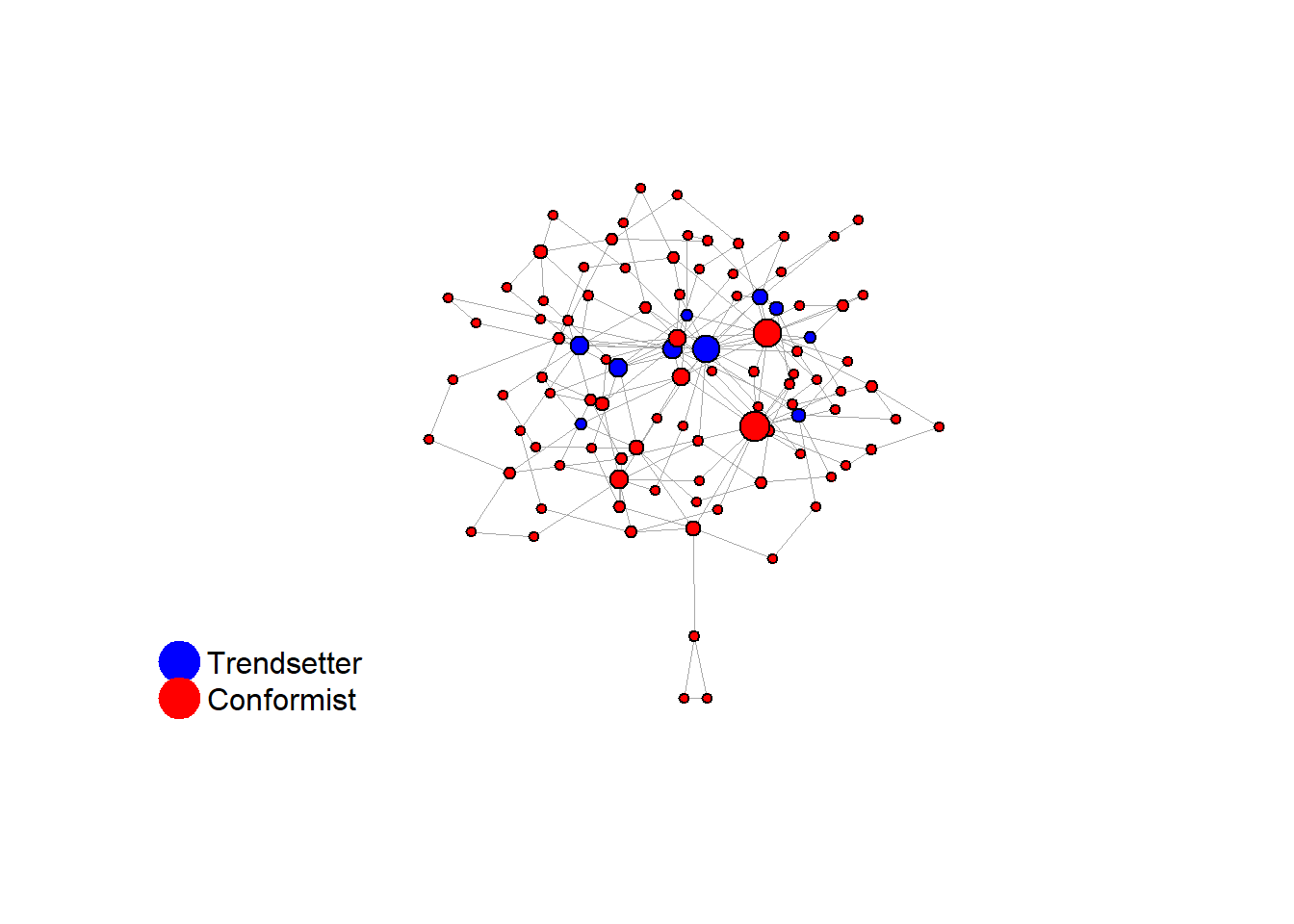
seed = seed + 1knitr::include_graphics(paste0("./figures/animation_network_", seed, ".gif"))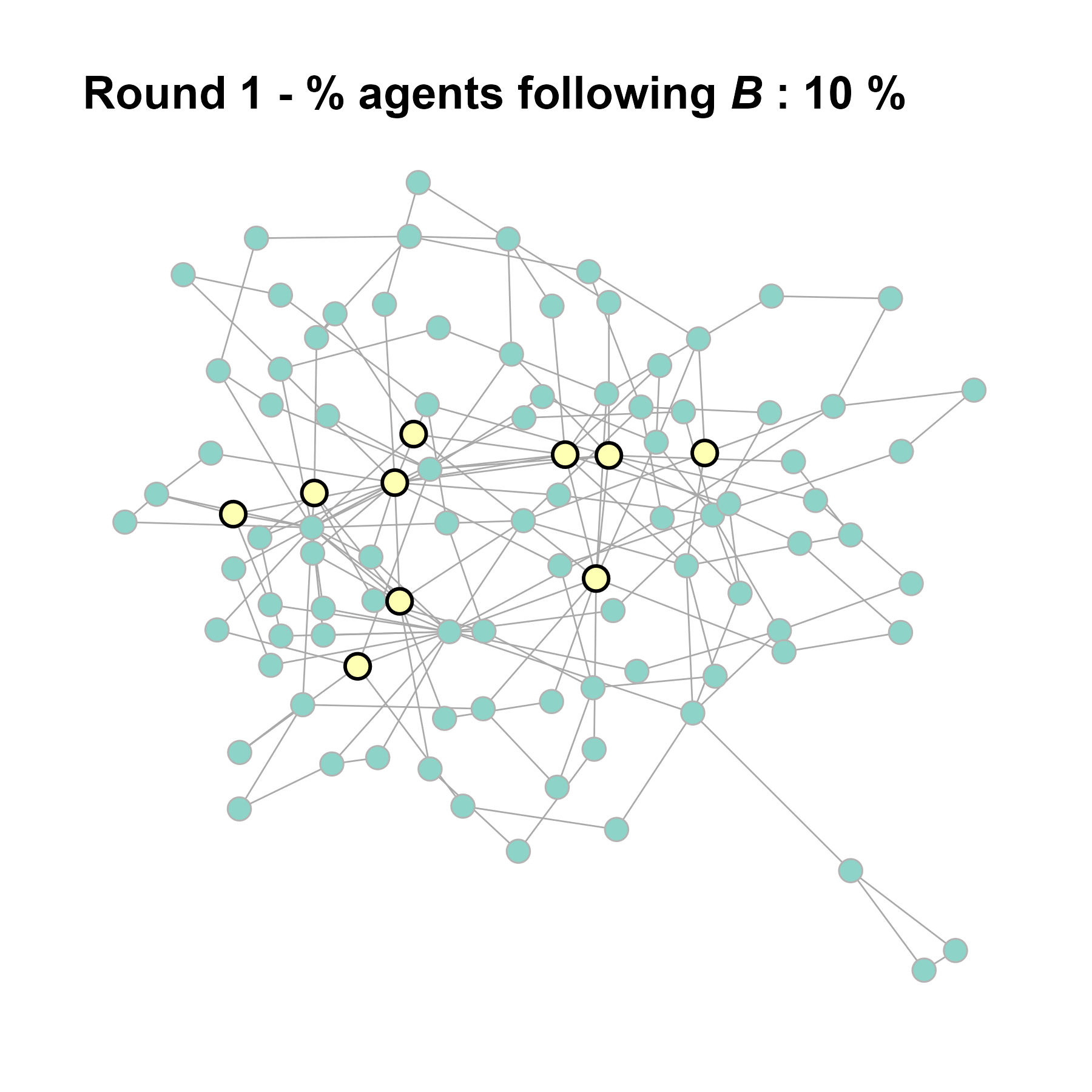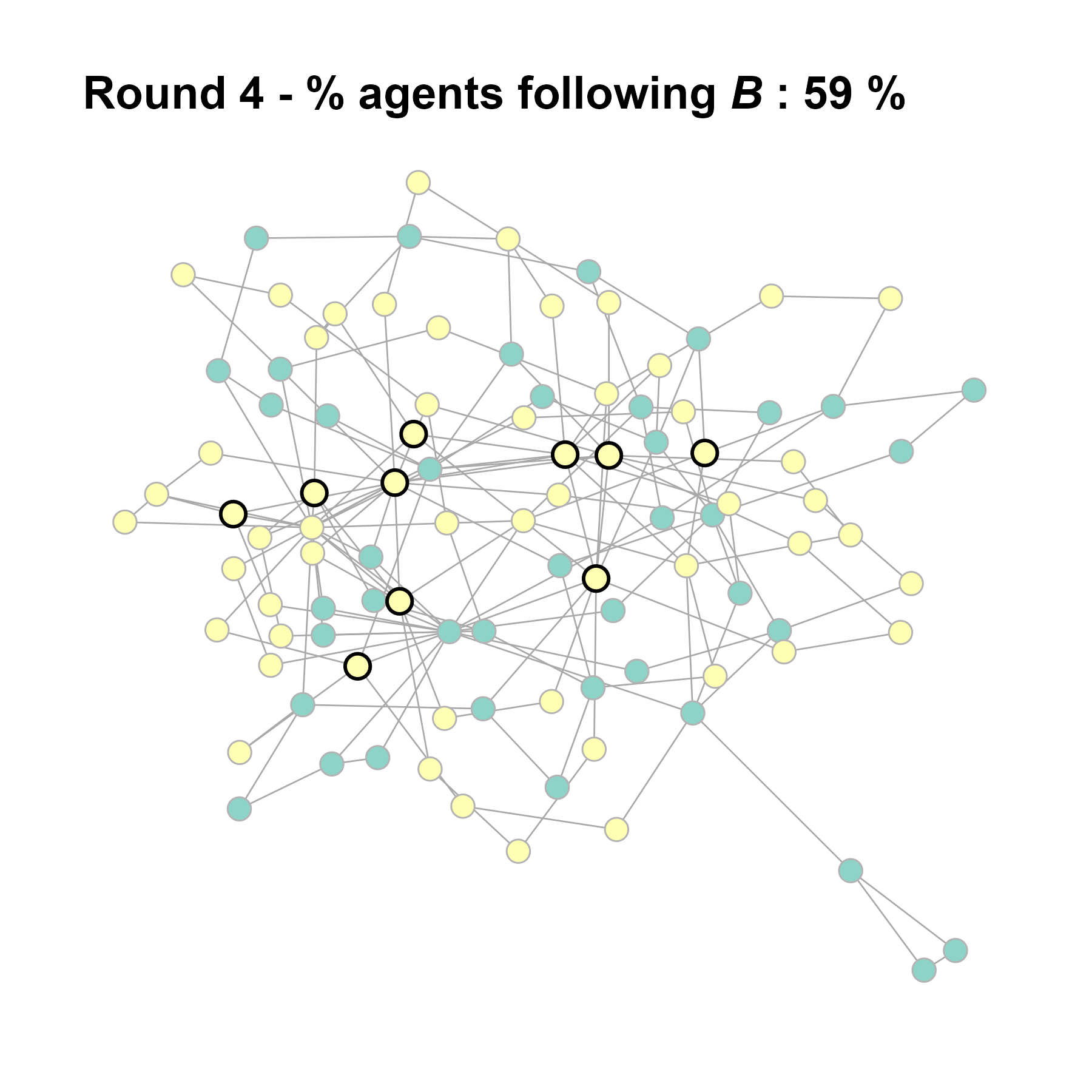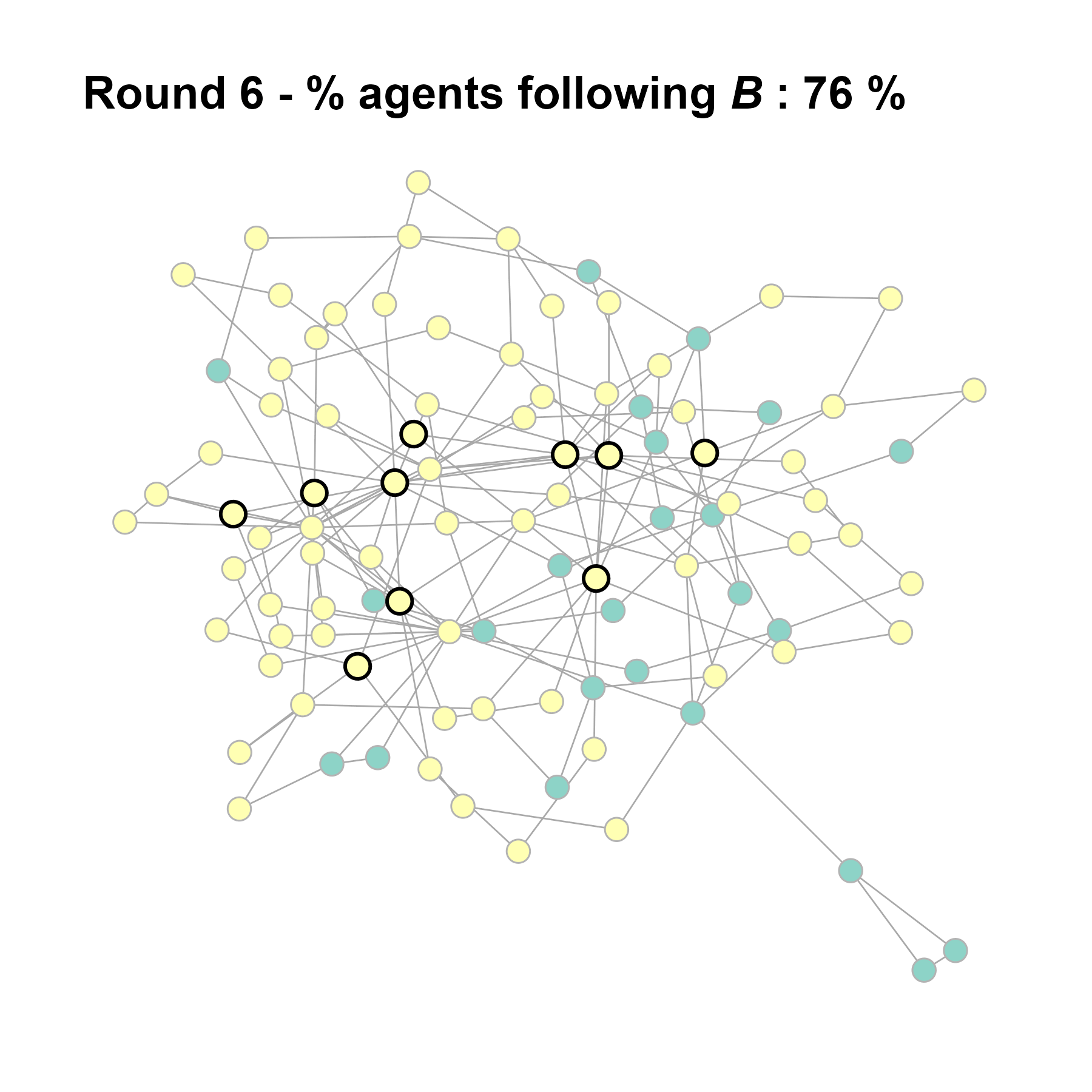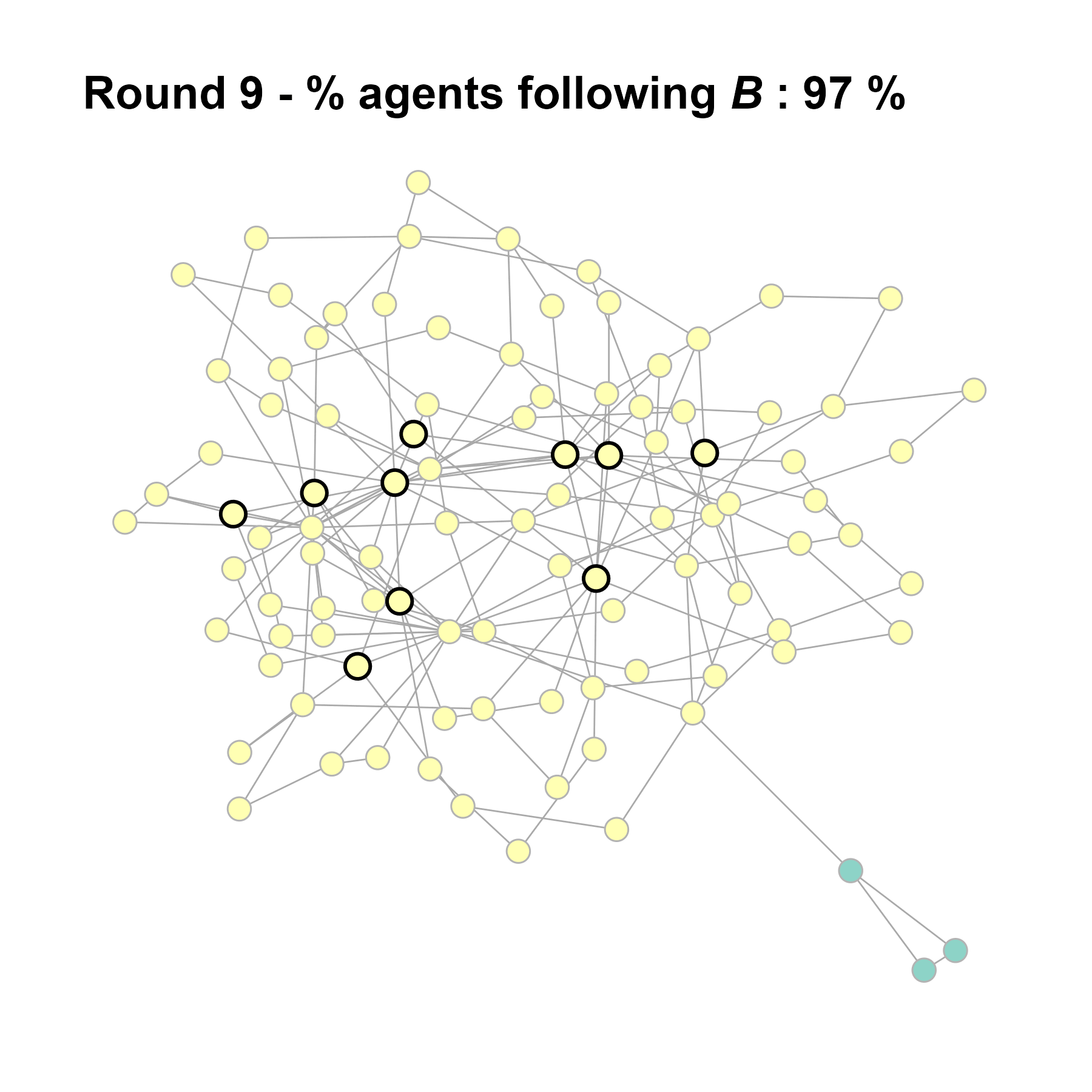
test <- run_one_seed(6, k_min = 2, k_max = 20, alpha = 2.4, rho = 0.4, r = -0.1)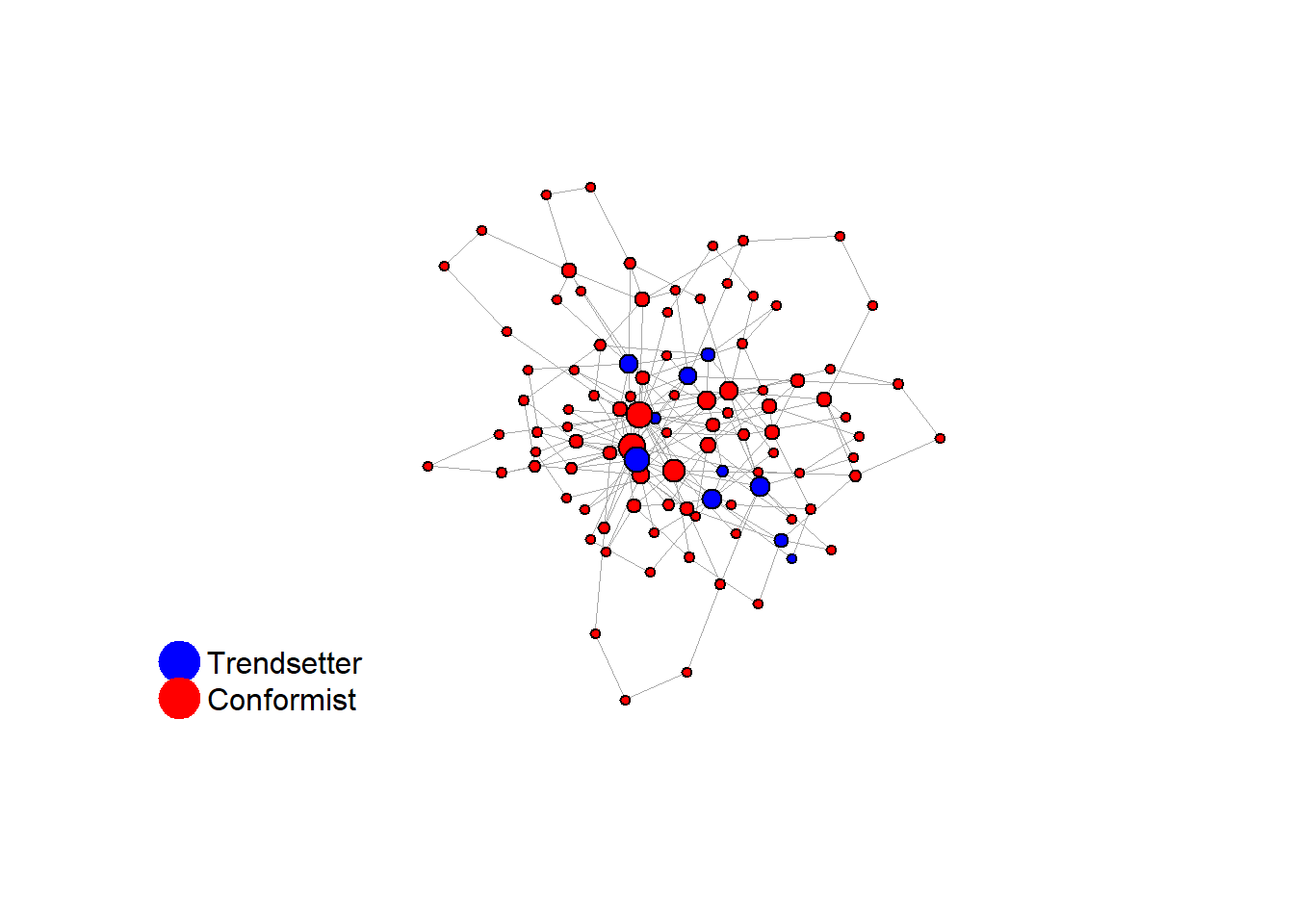
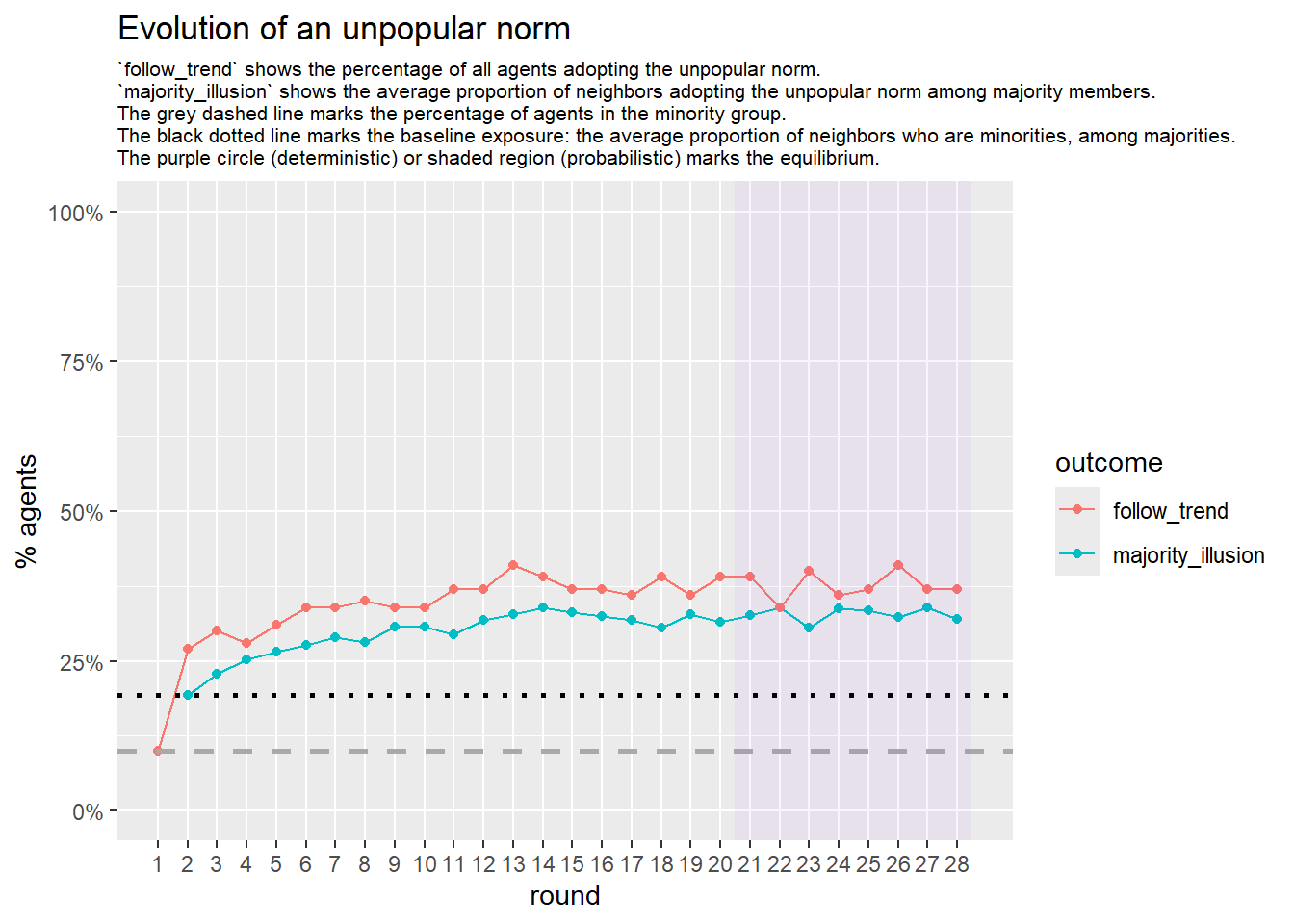
seed = seed + 1knitr::include_graphics(paste0("./figures/animation_network_", seed, ".gif"))
test <- run_one_seed(7, k_min = 2, k_max = 20, alpha = 2.4, rho = 0.4, r = -0.1)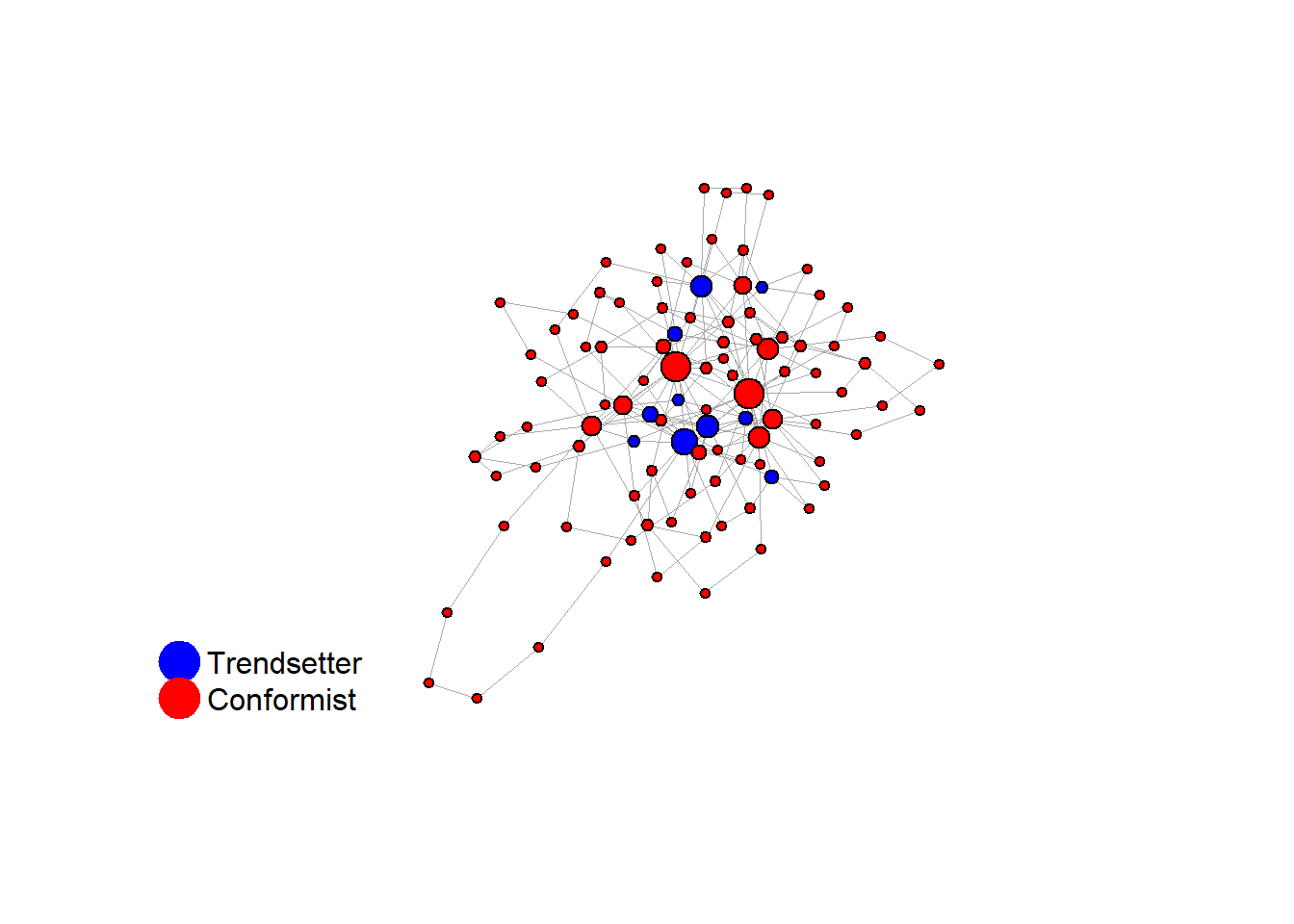
seed = seed + 1knitr::include_graphics(paste0("./figures/animation_network_", seed, ".gif"))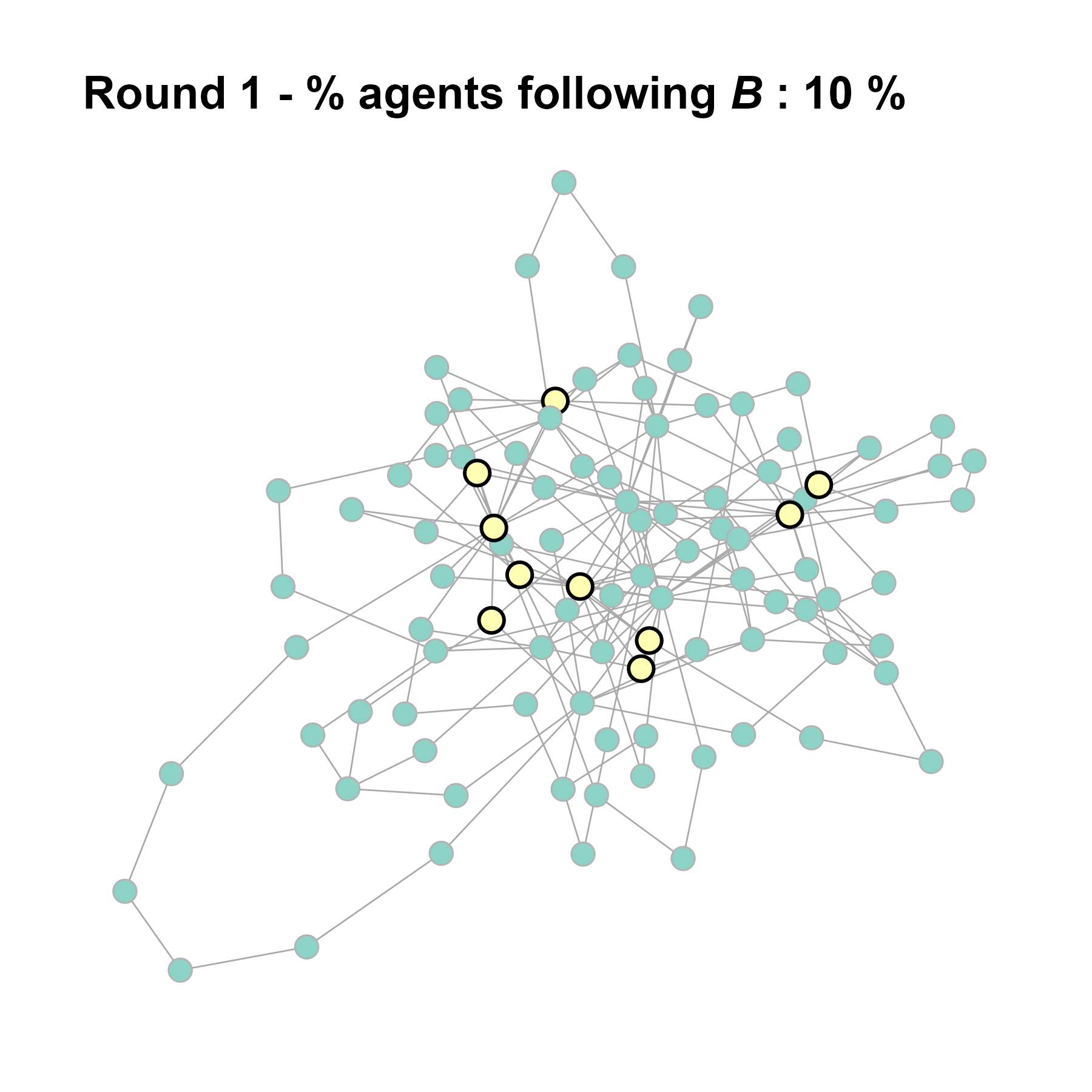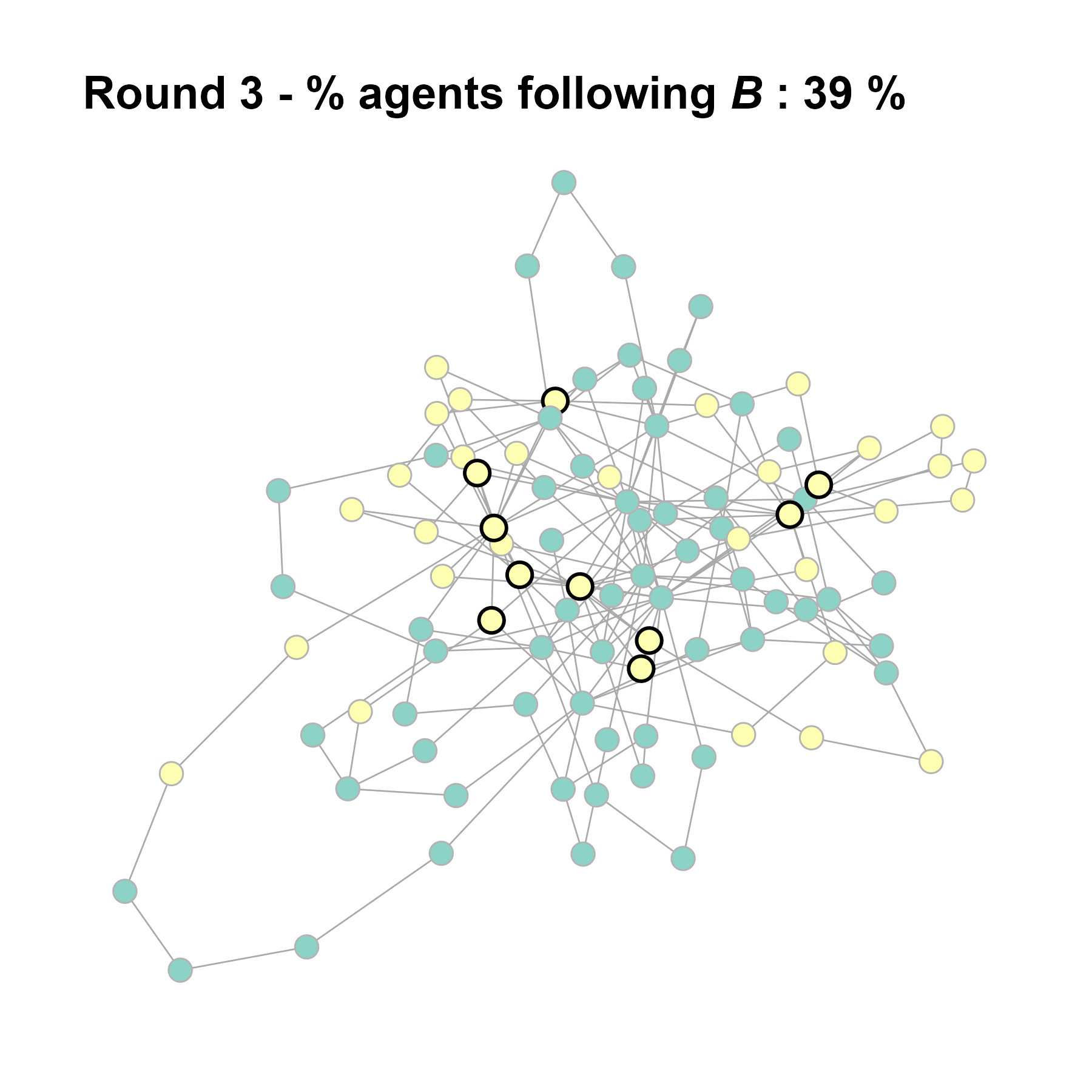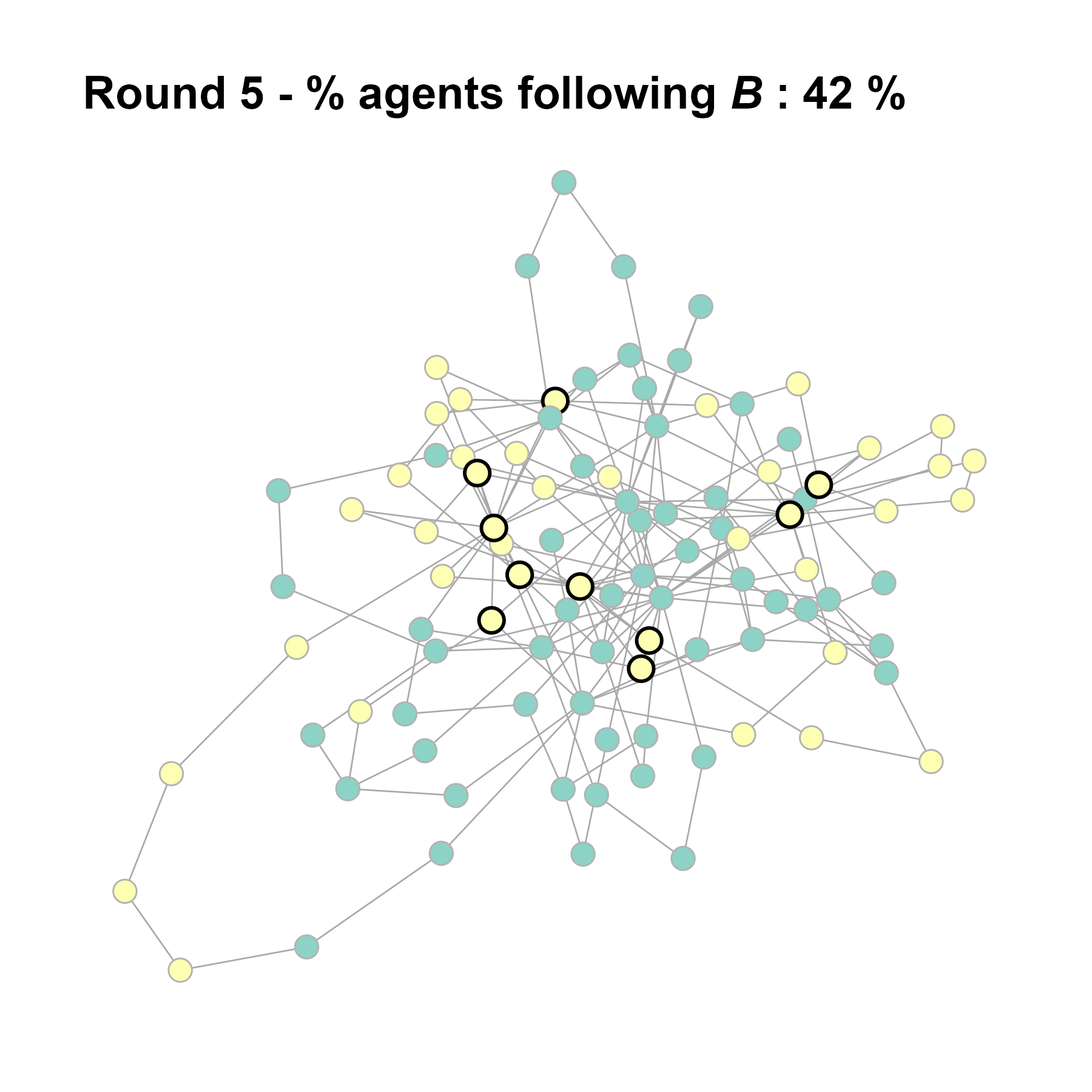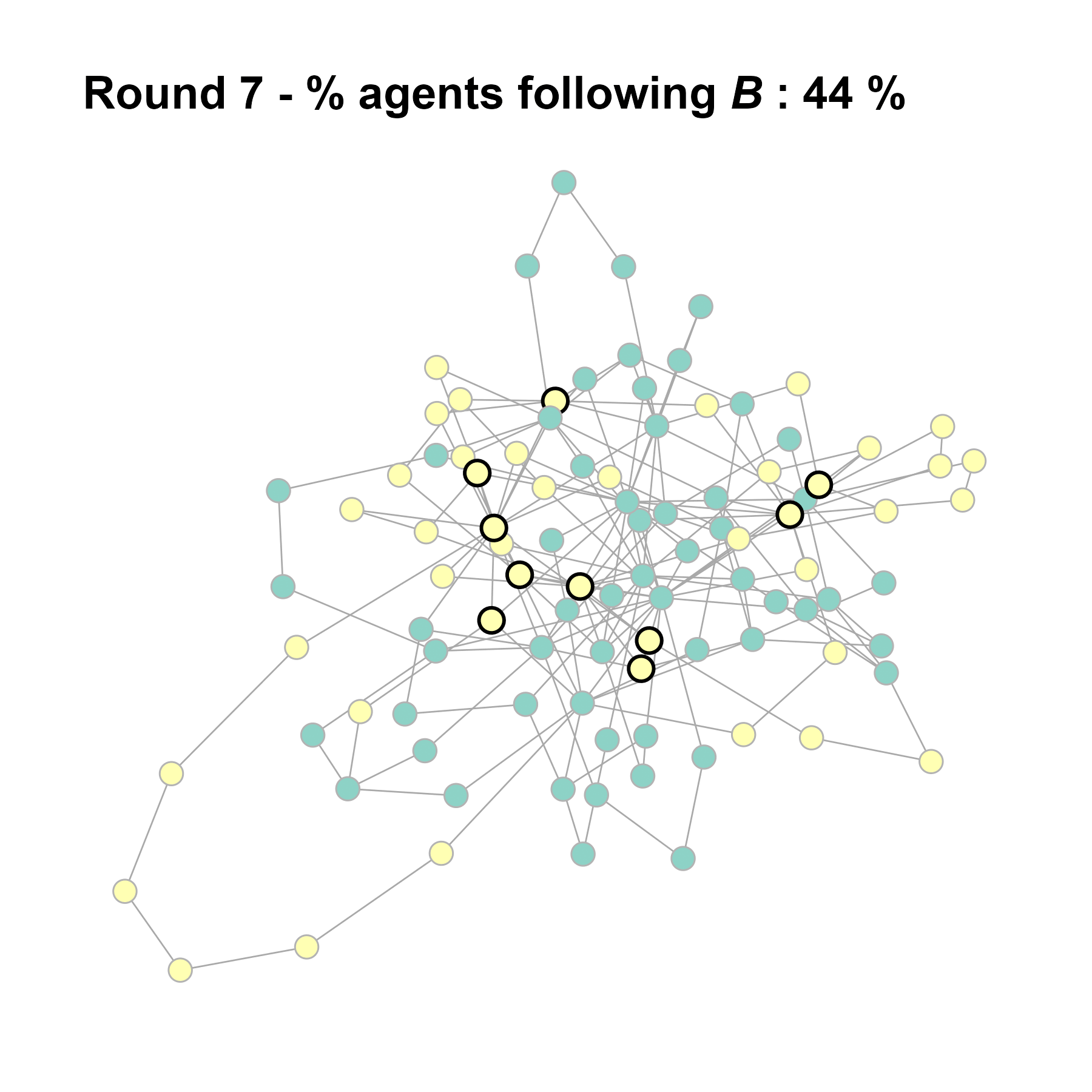
Increase the density:
test <- run_one_seed(8, k_min = 4, k_max = 99, alpha = 2.1, rho = 0.5, r = -0.4)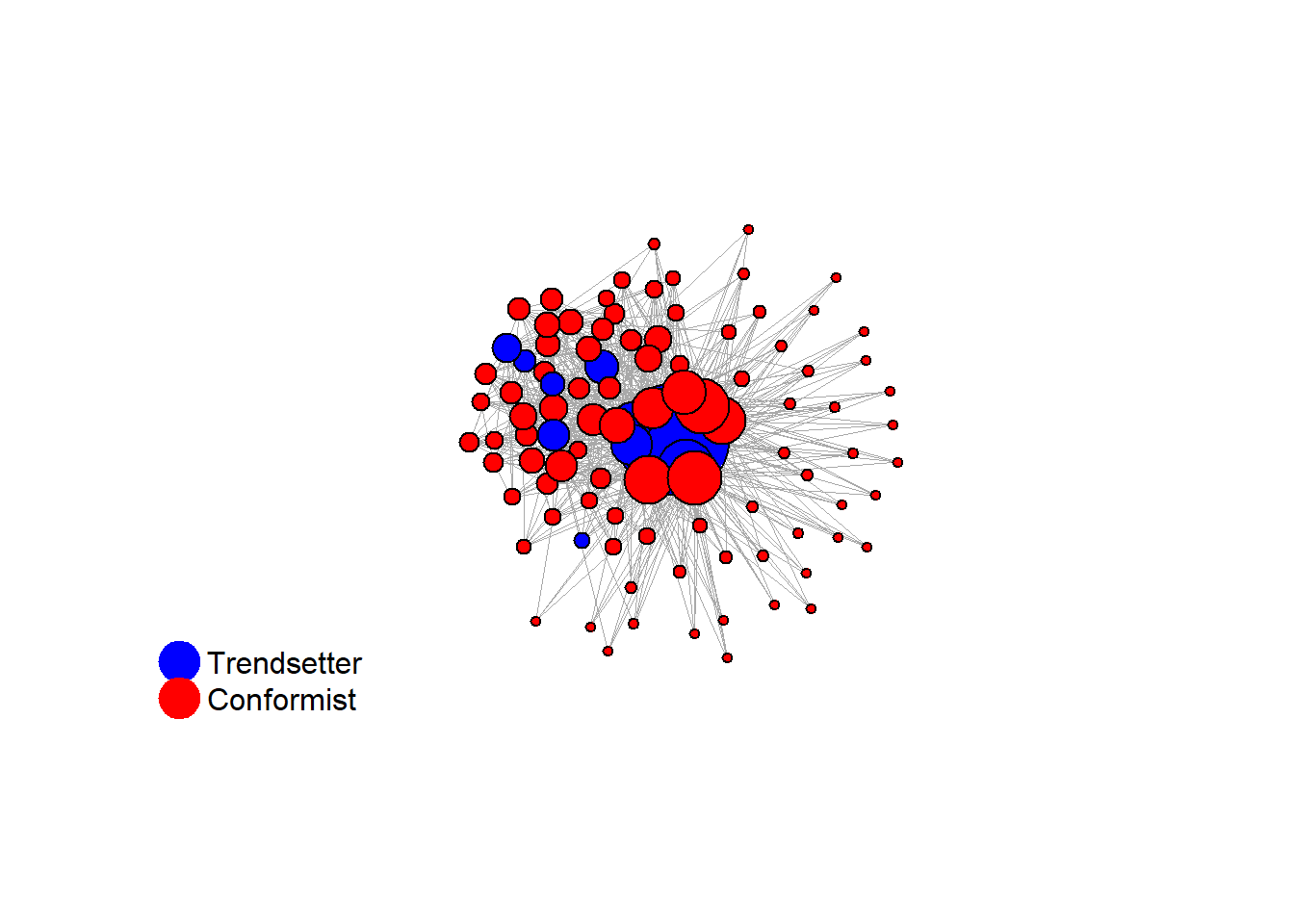
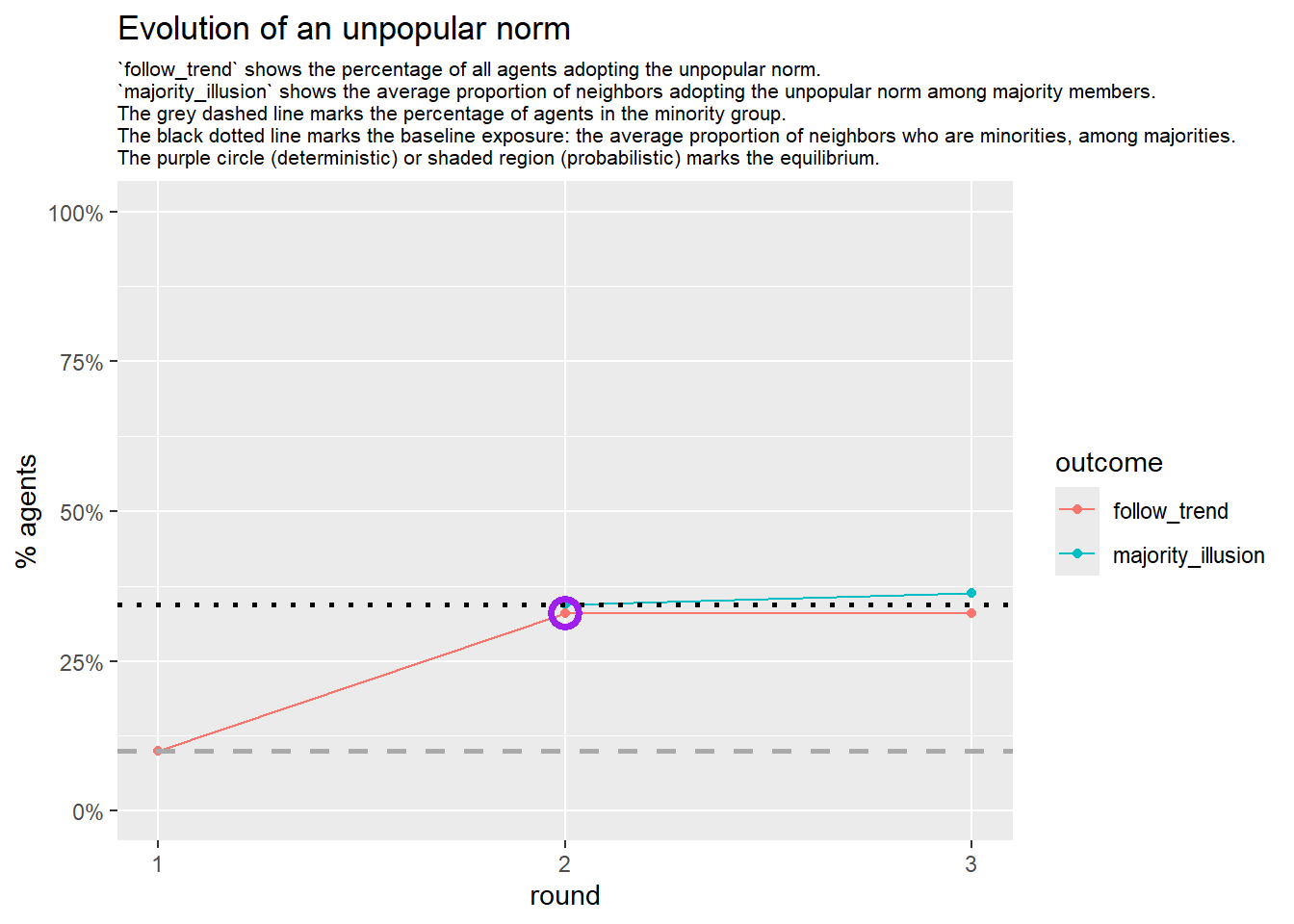
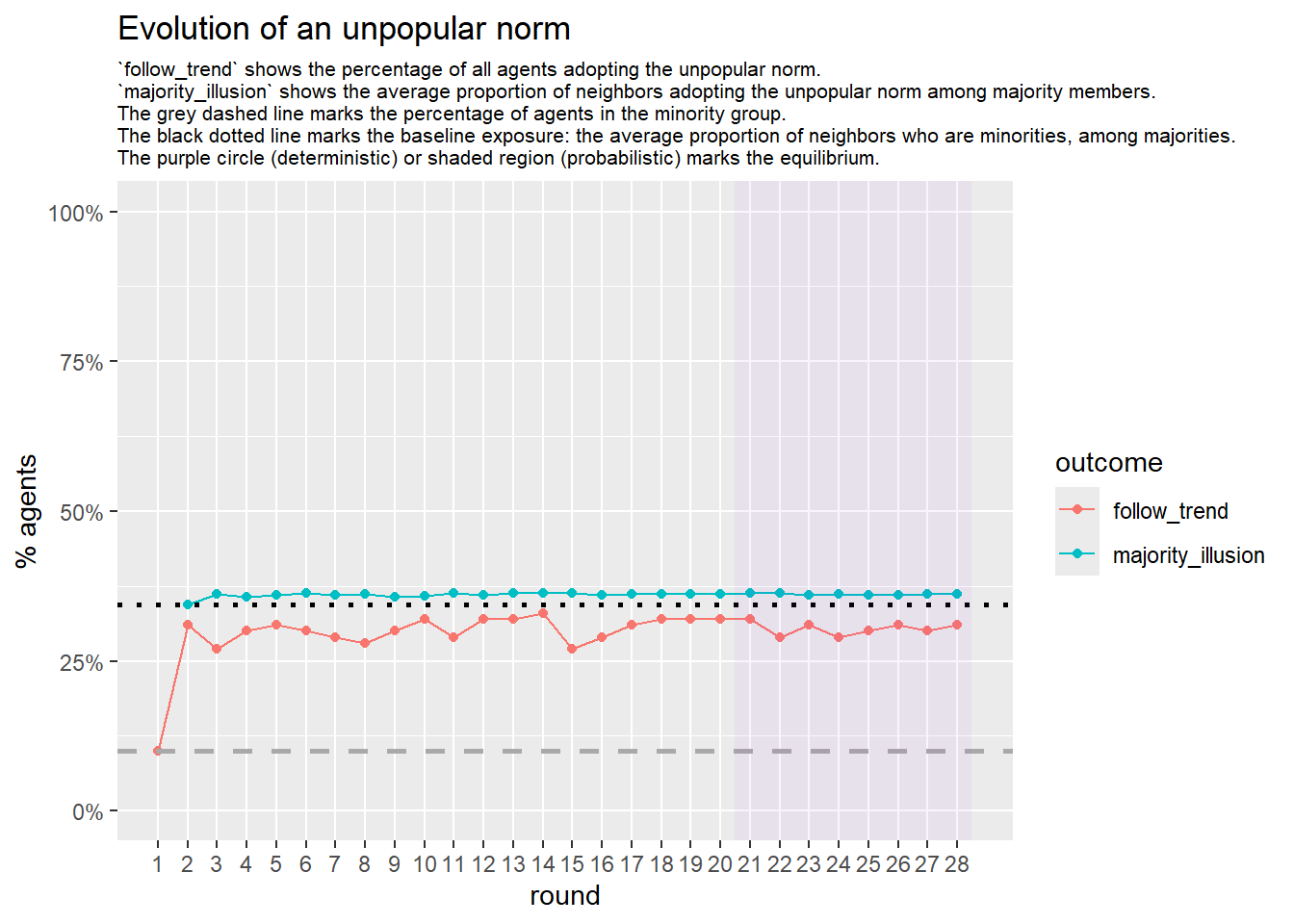
seed = seed + 1knitr::include_graphics(paste0("./figures/animation_network_", seed, ".gif"))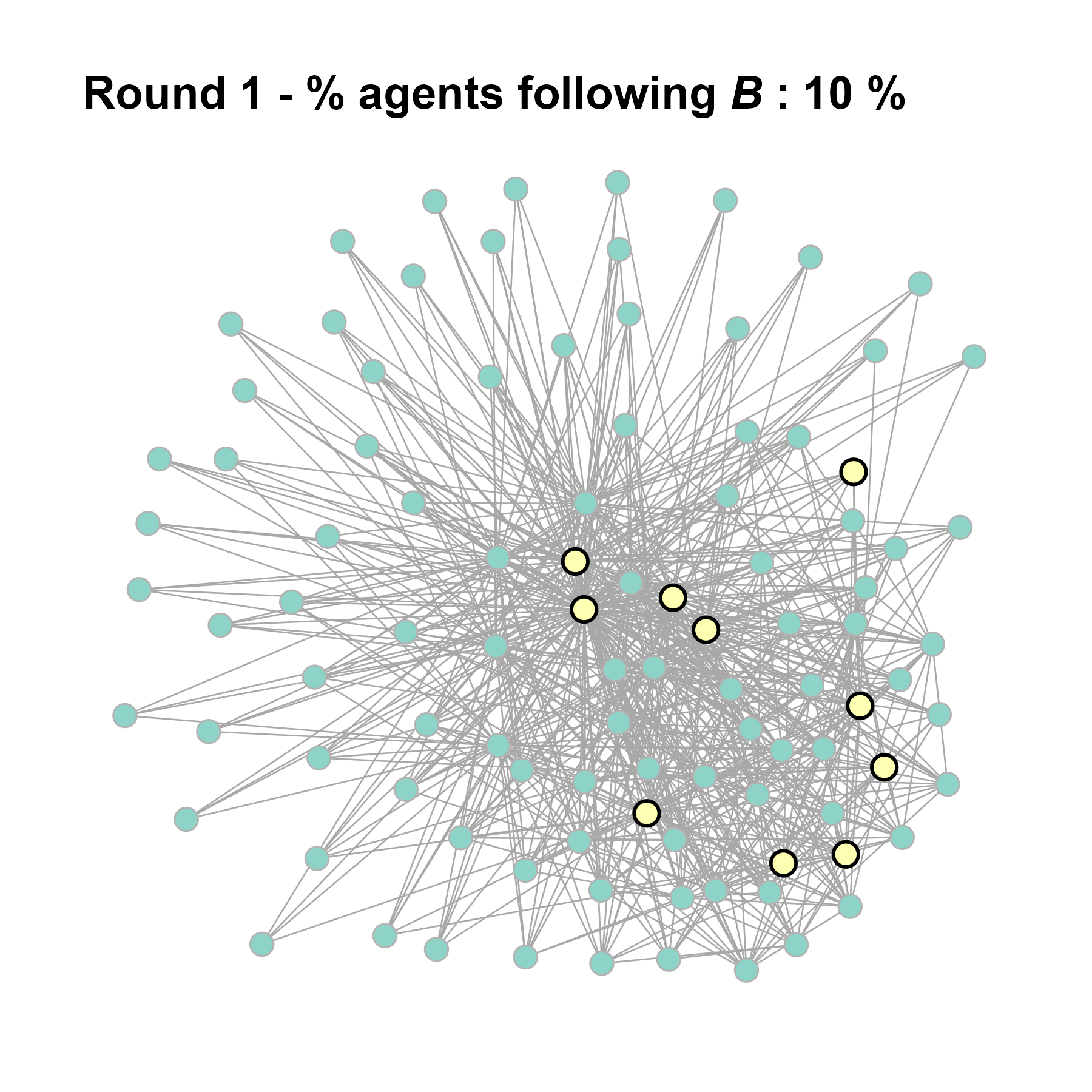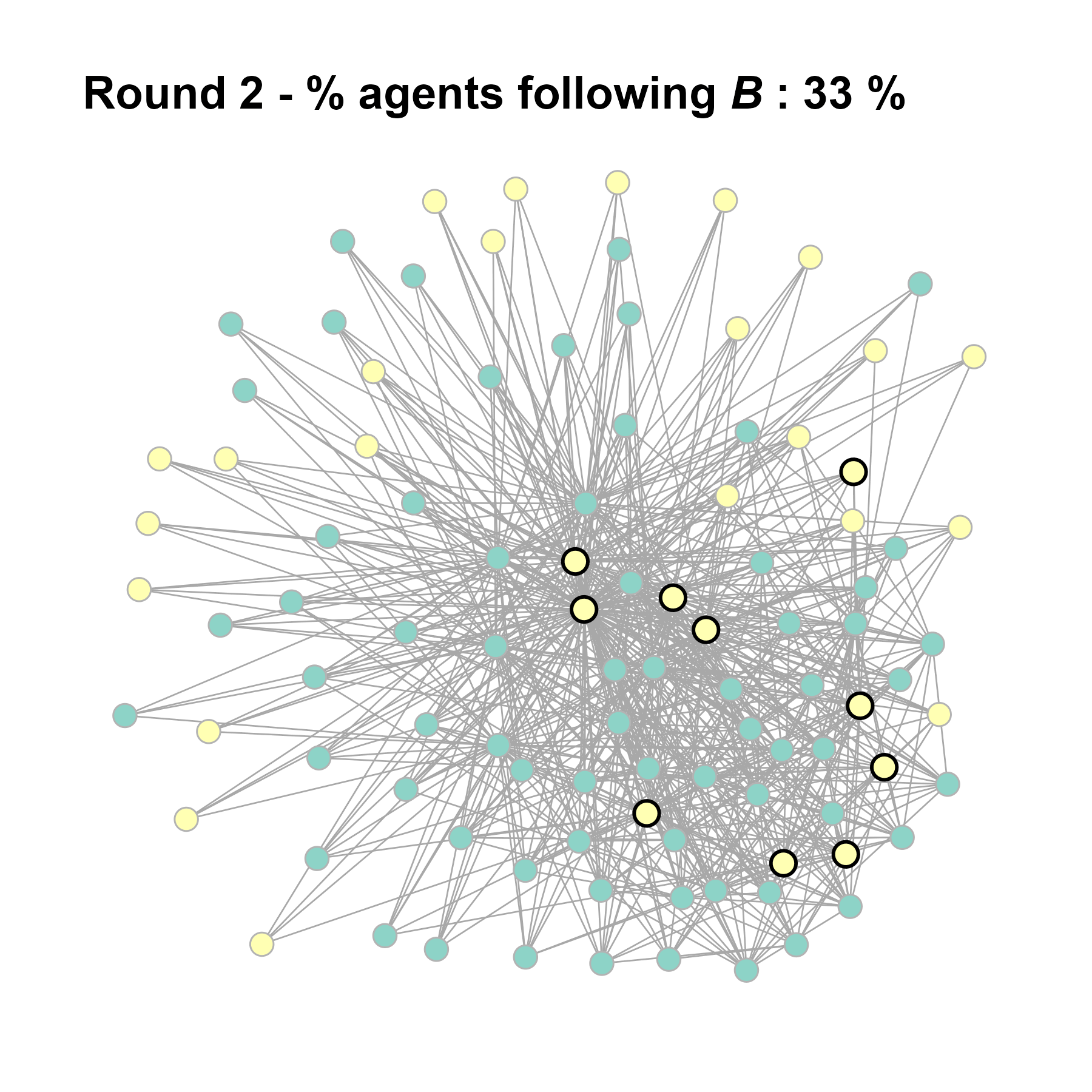
test <- run_one_seed(9, k_min = 4, k_max = 99, alpha = 2.1, rho = 0.5, r = -0.4)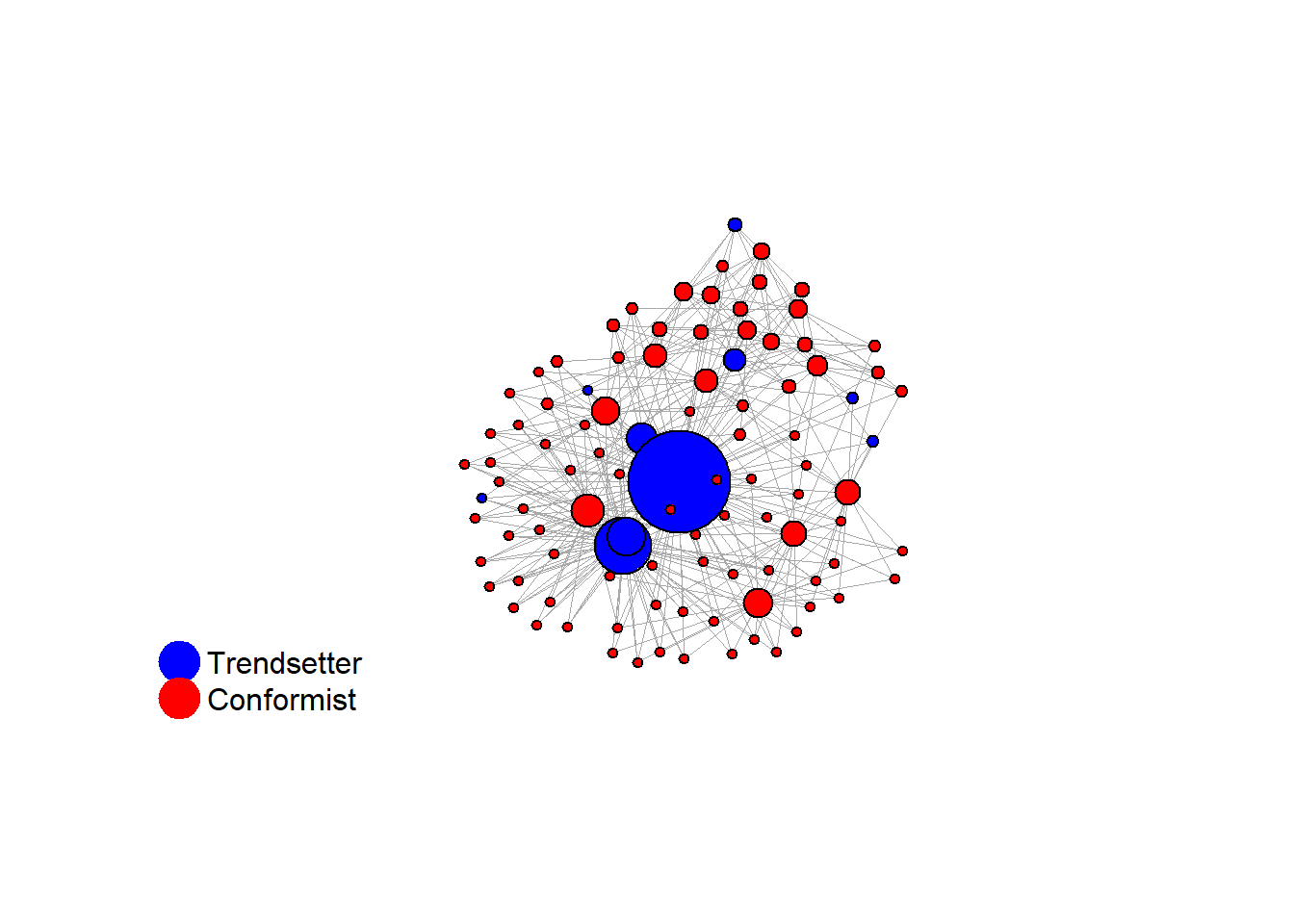
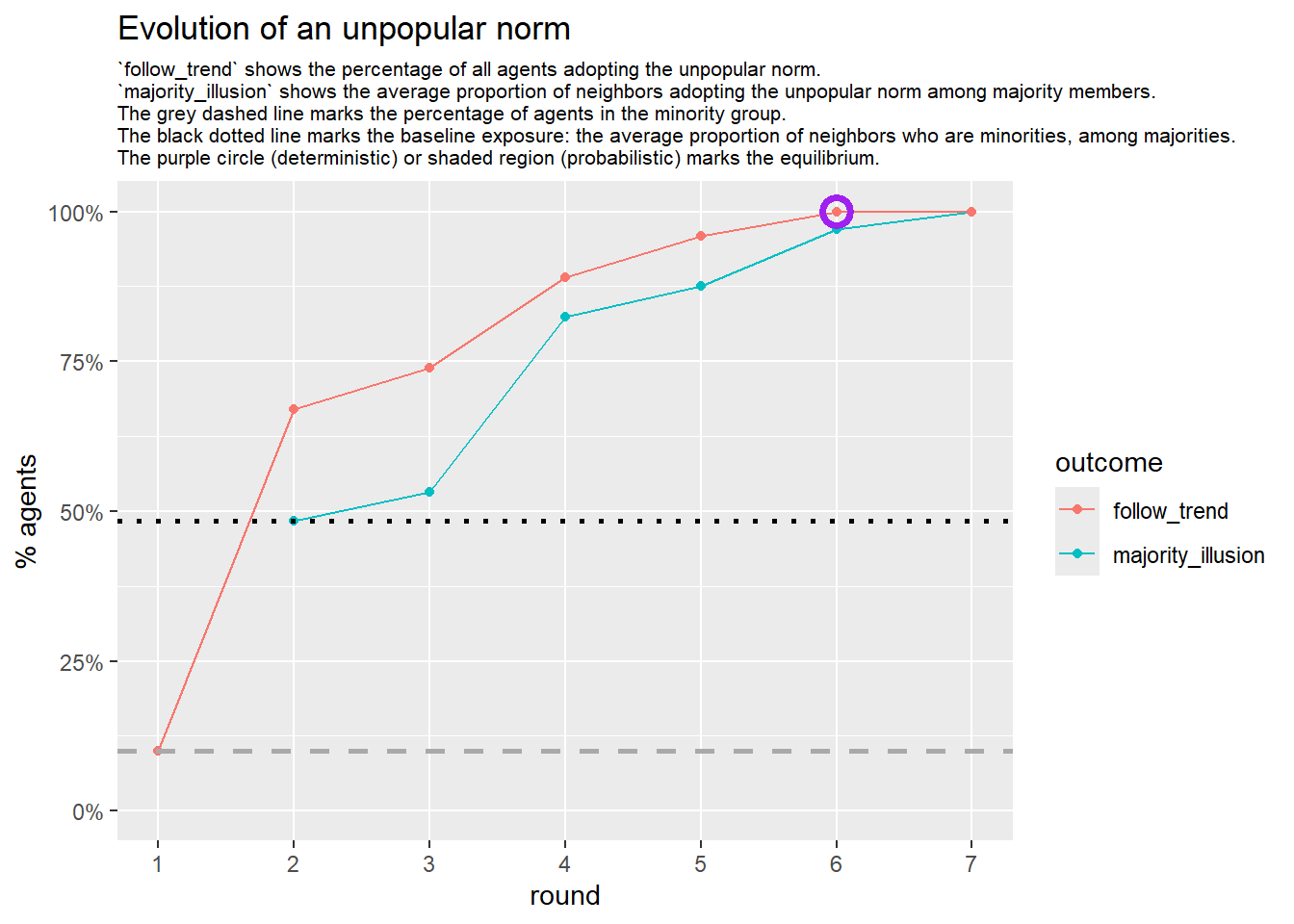
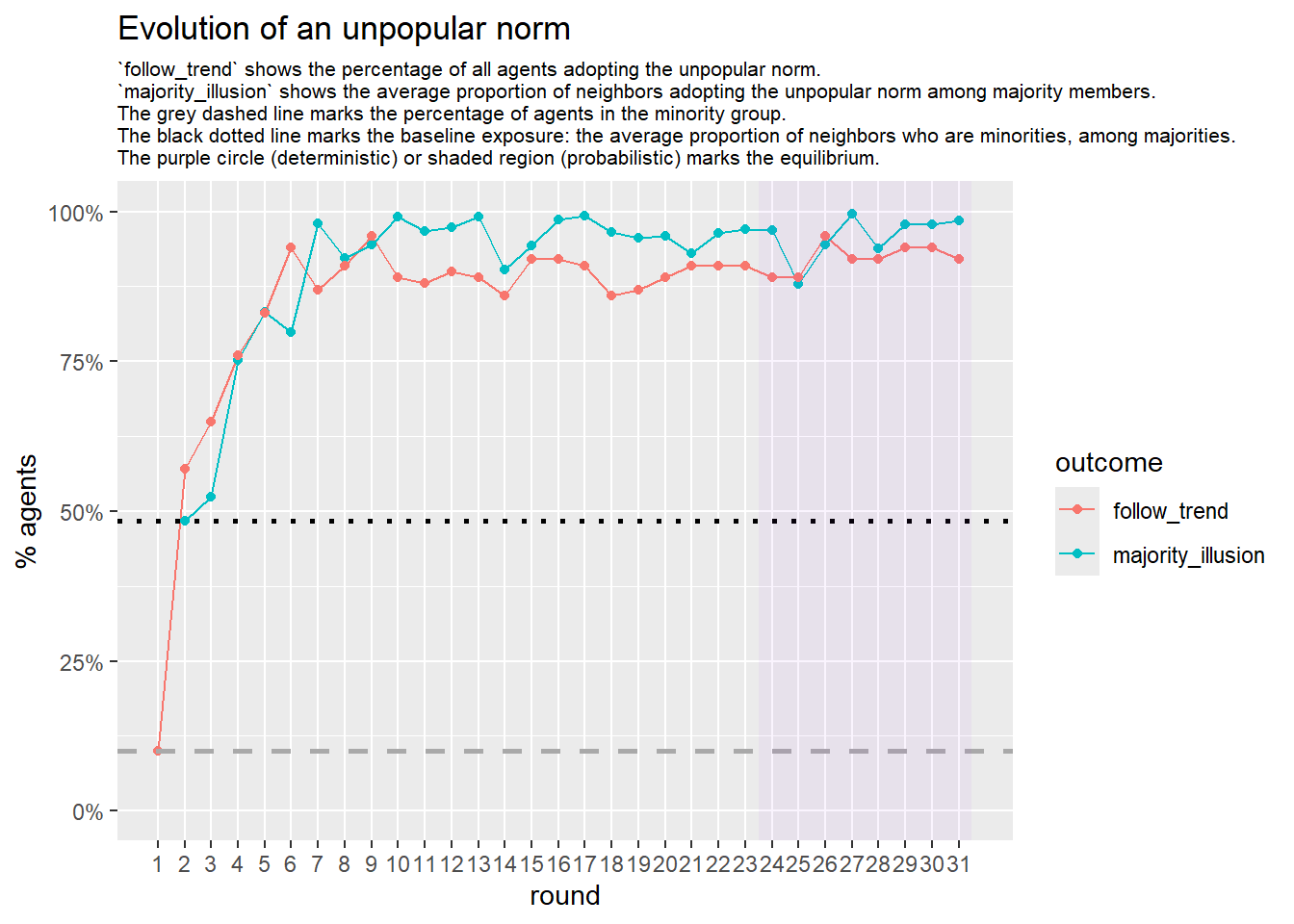
seed = seed + 1knitr::include_graphics(paste0("./figures/animation_network_", seed, ".gif"))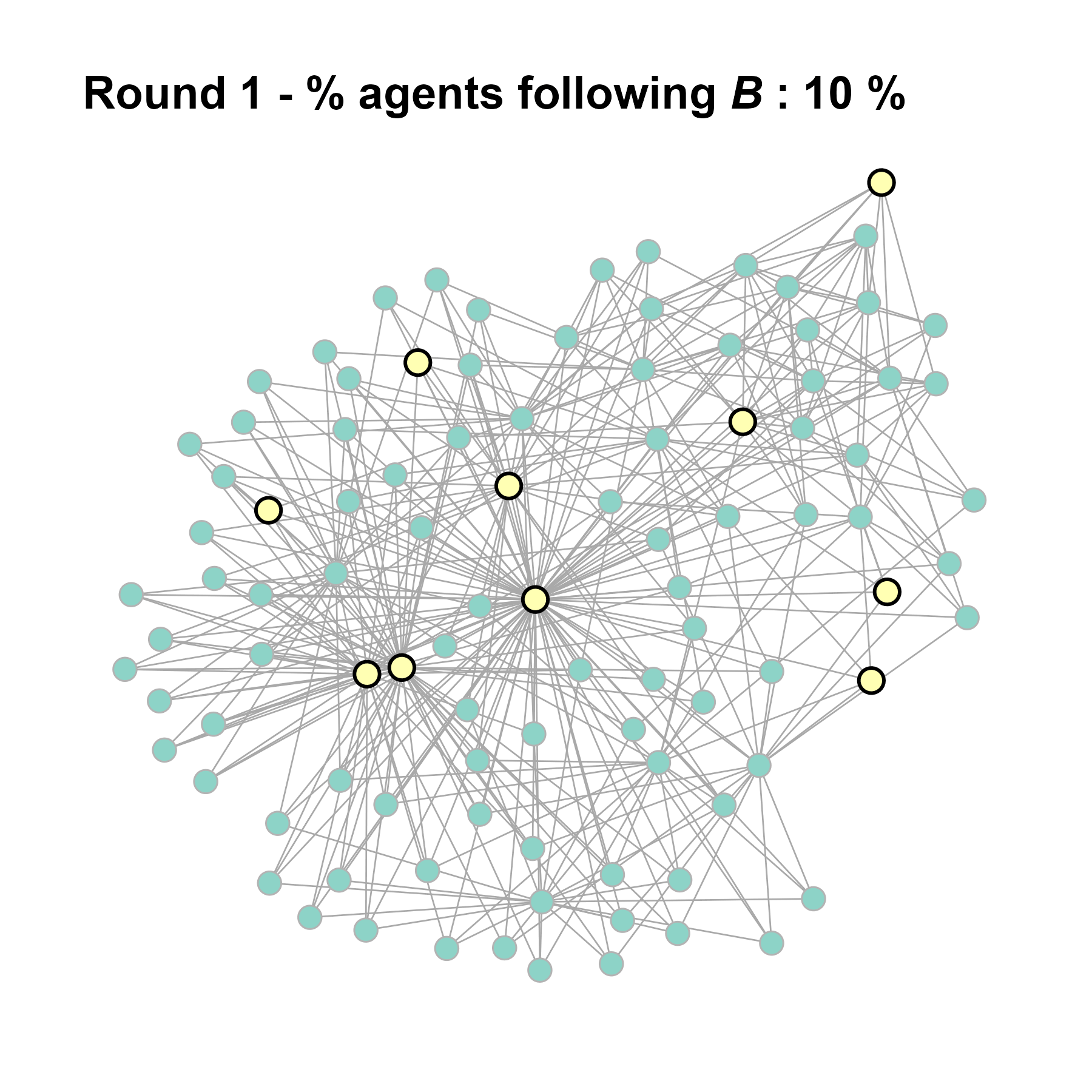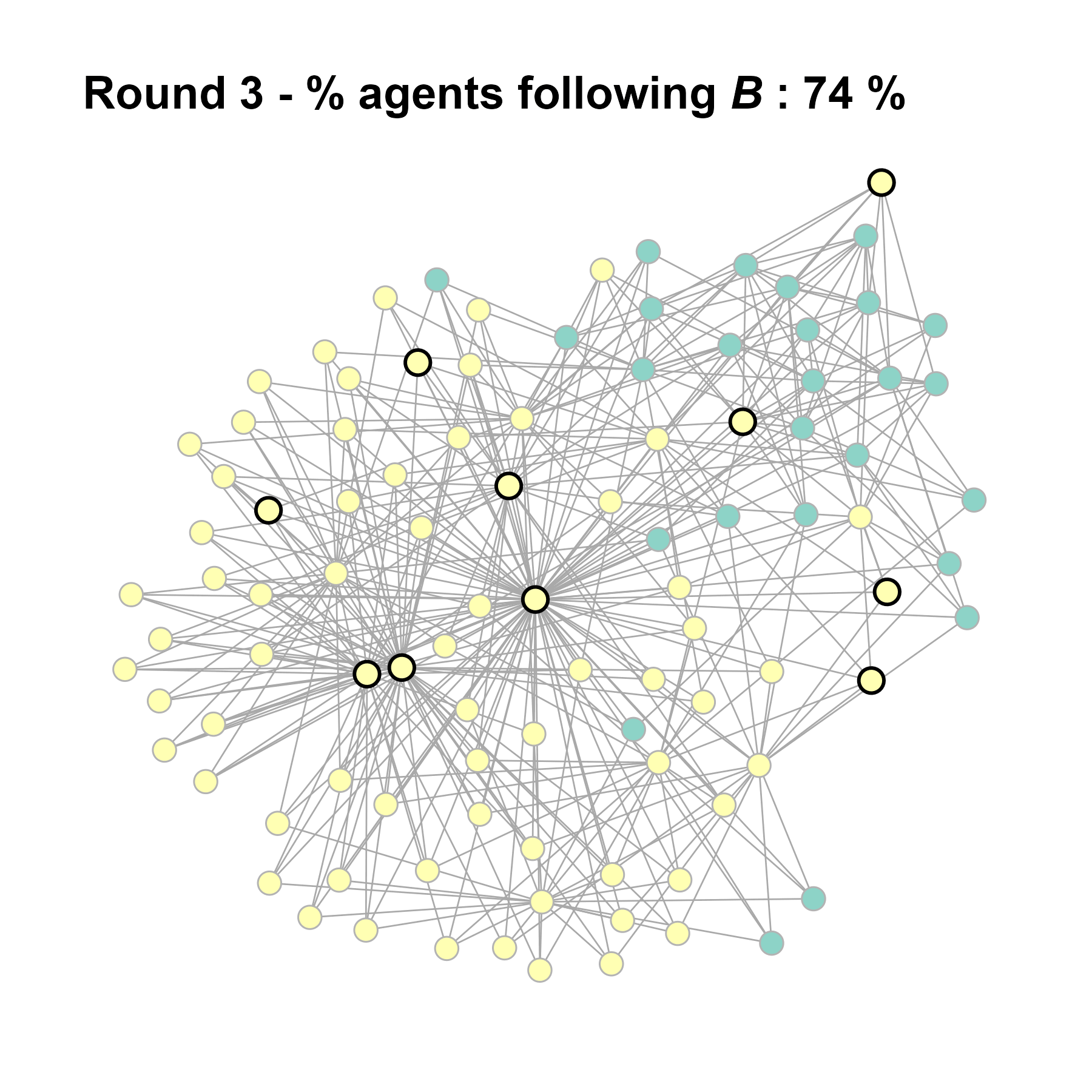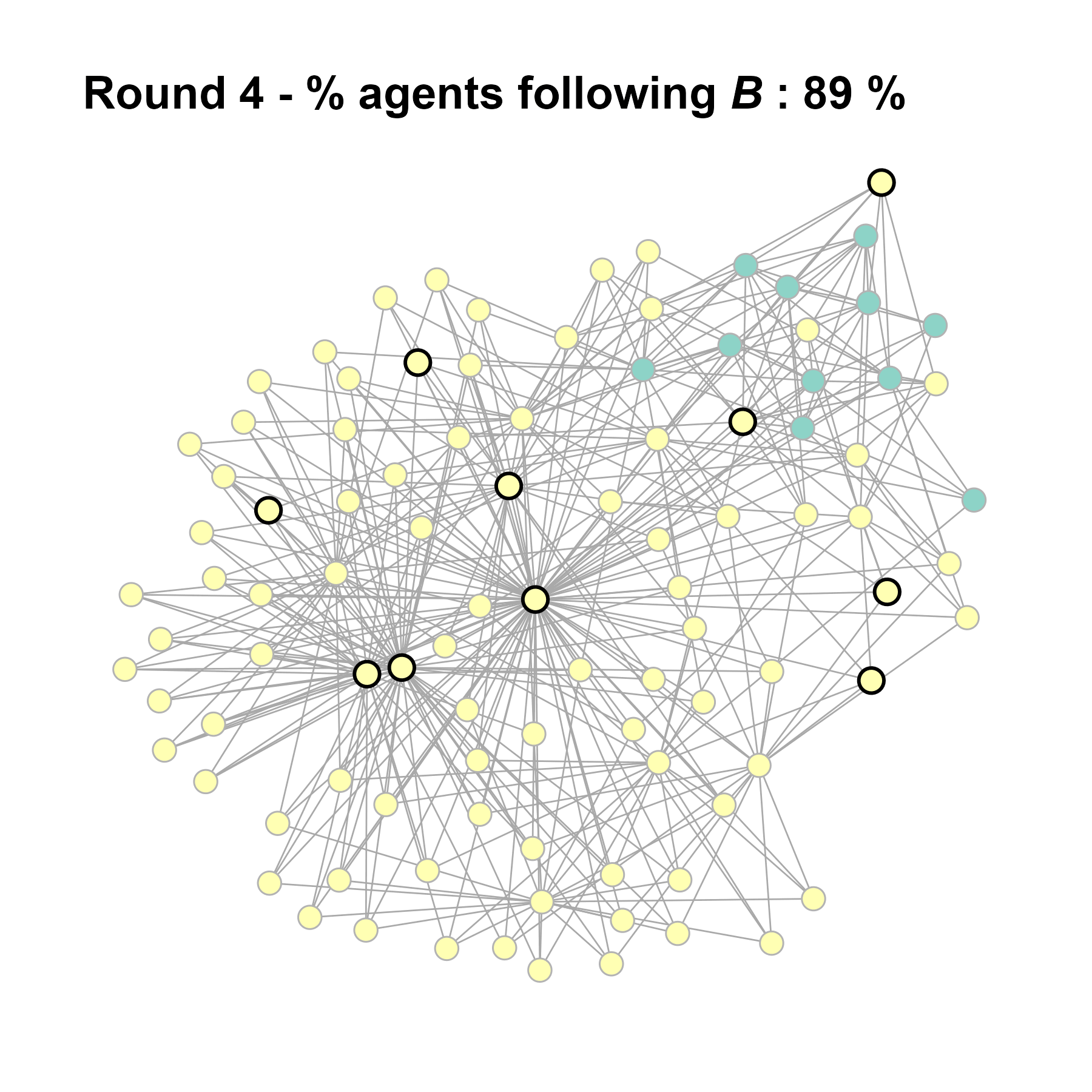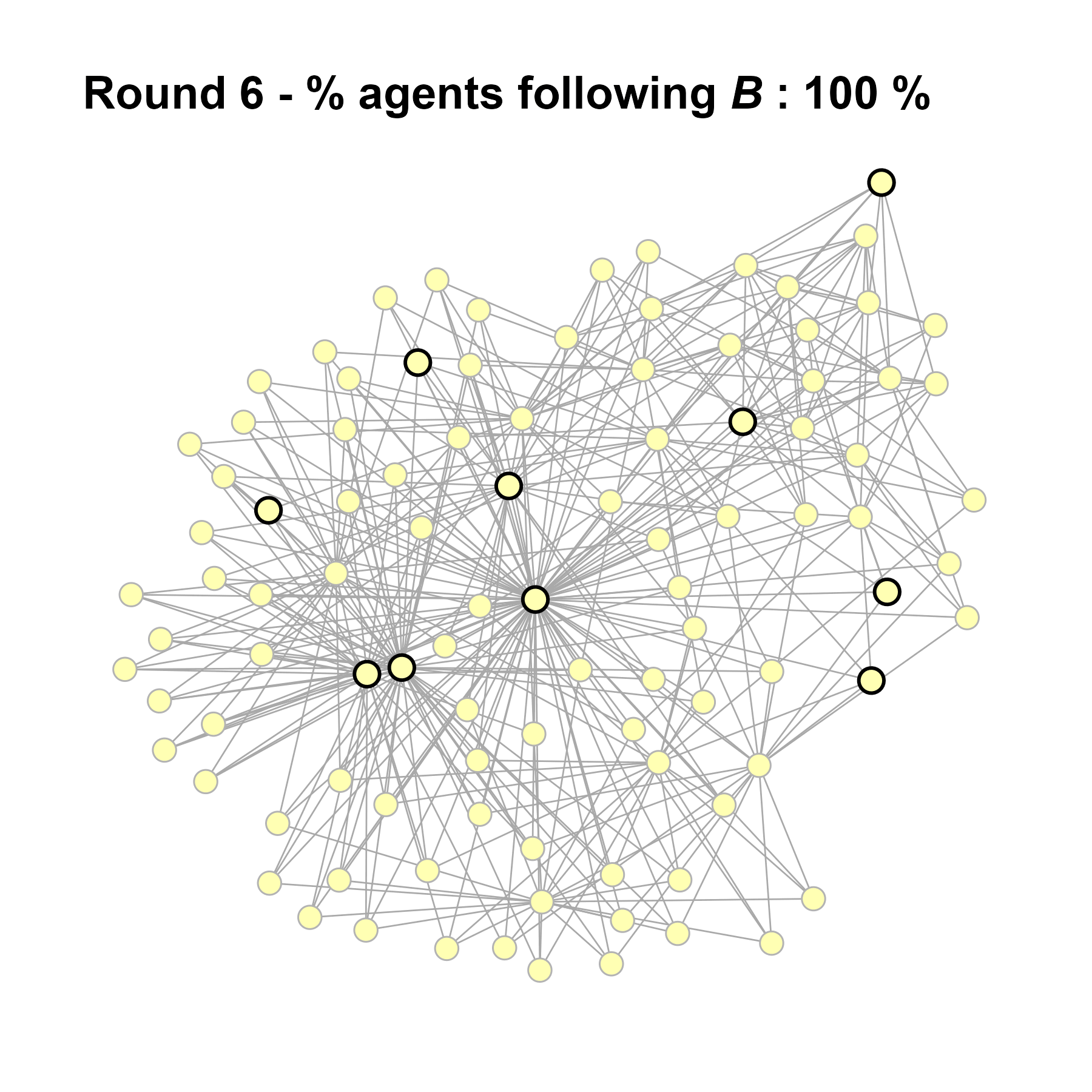
test <- run_one_seed(10, k_min = 4, k_max = 99, alpha = 2.1, rho = 0.5, r = -0.4)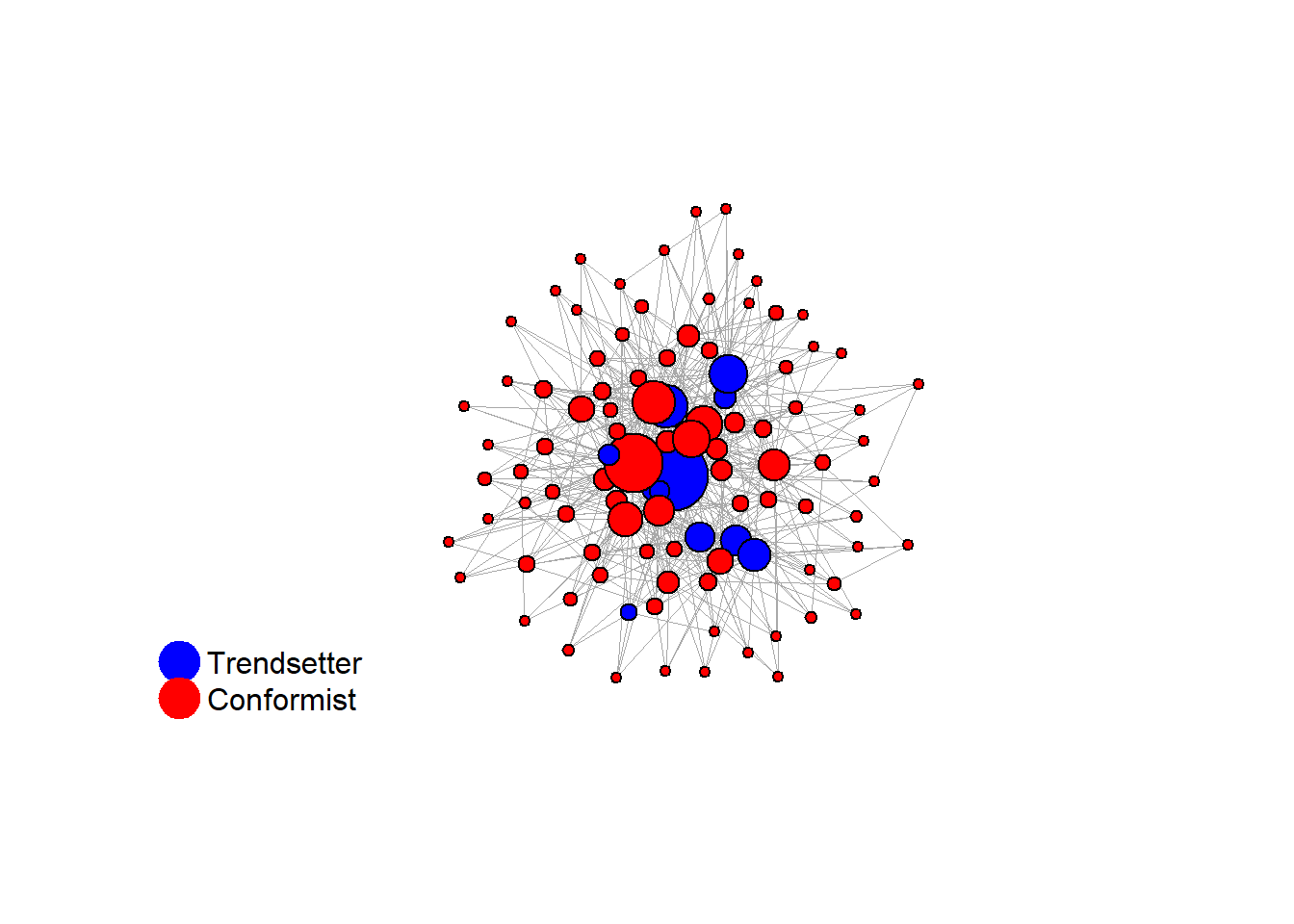
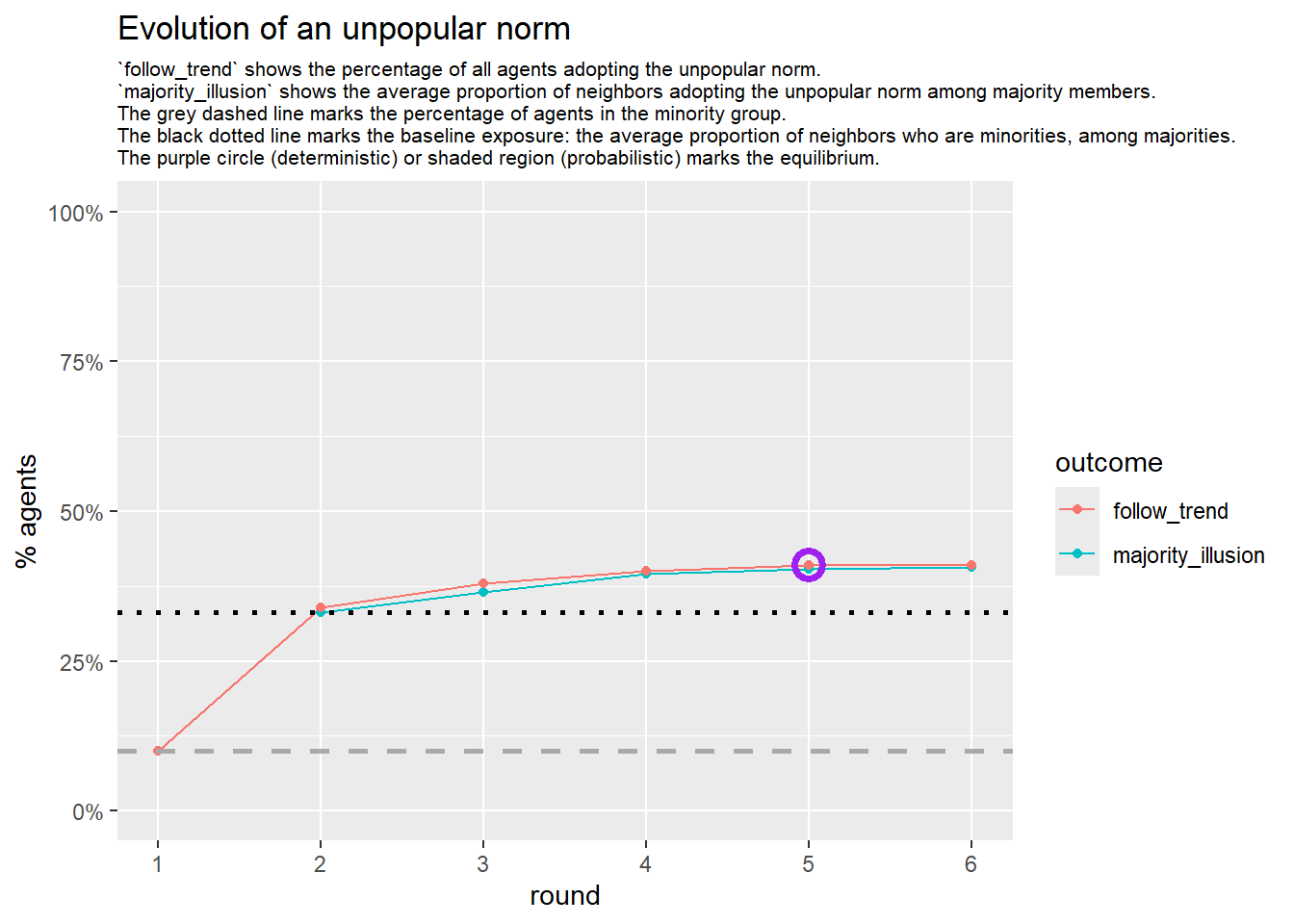
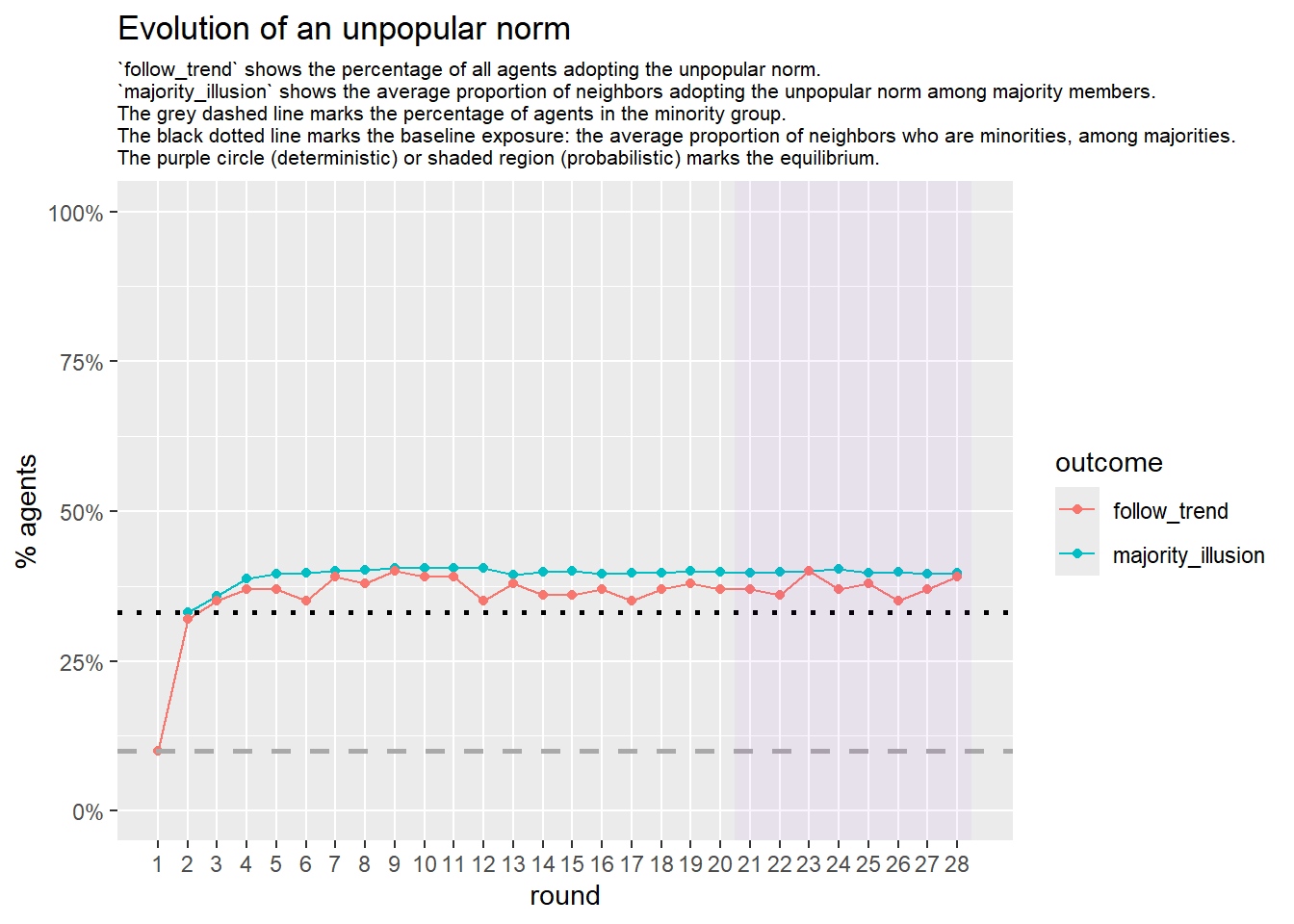
seed = seed + 1knitr::include_graphics(paste0("./figures/animation_network_", seed, ".gif"))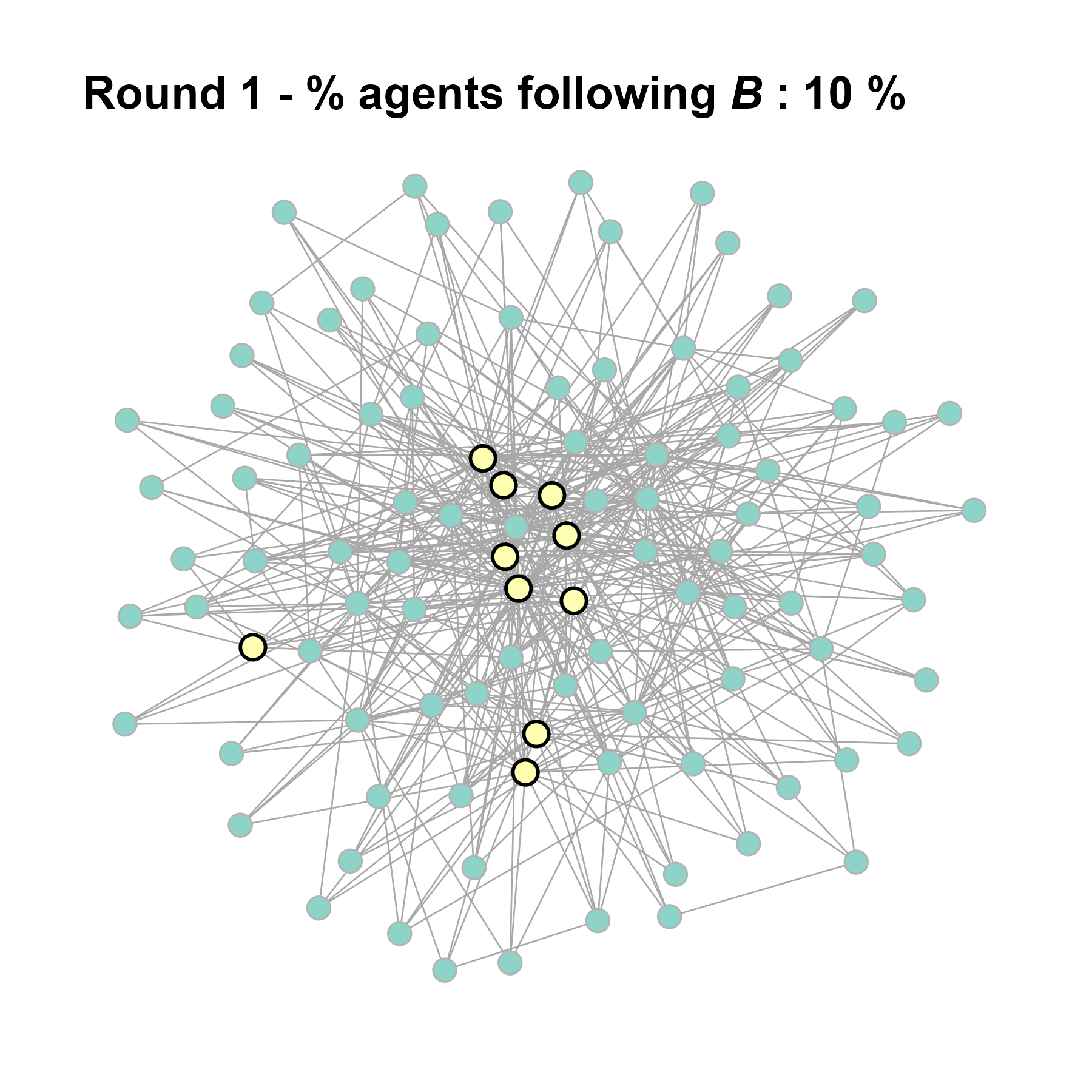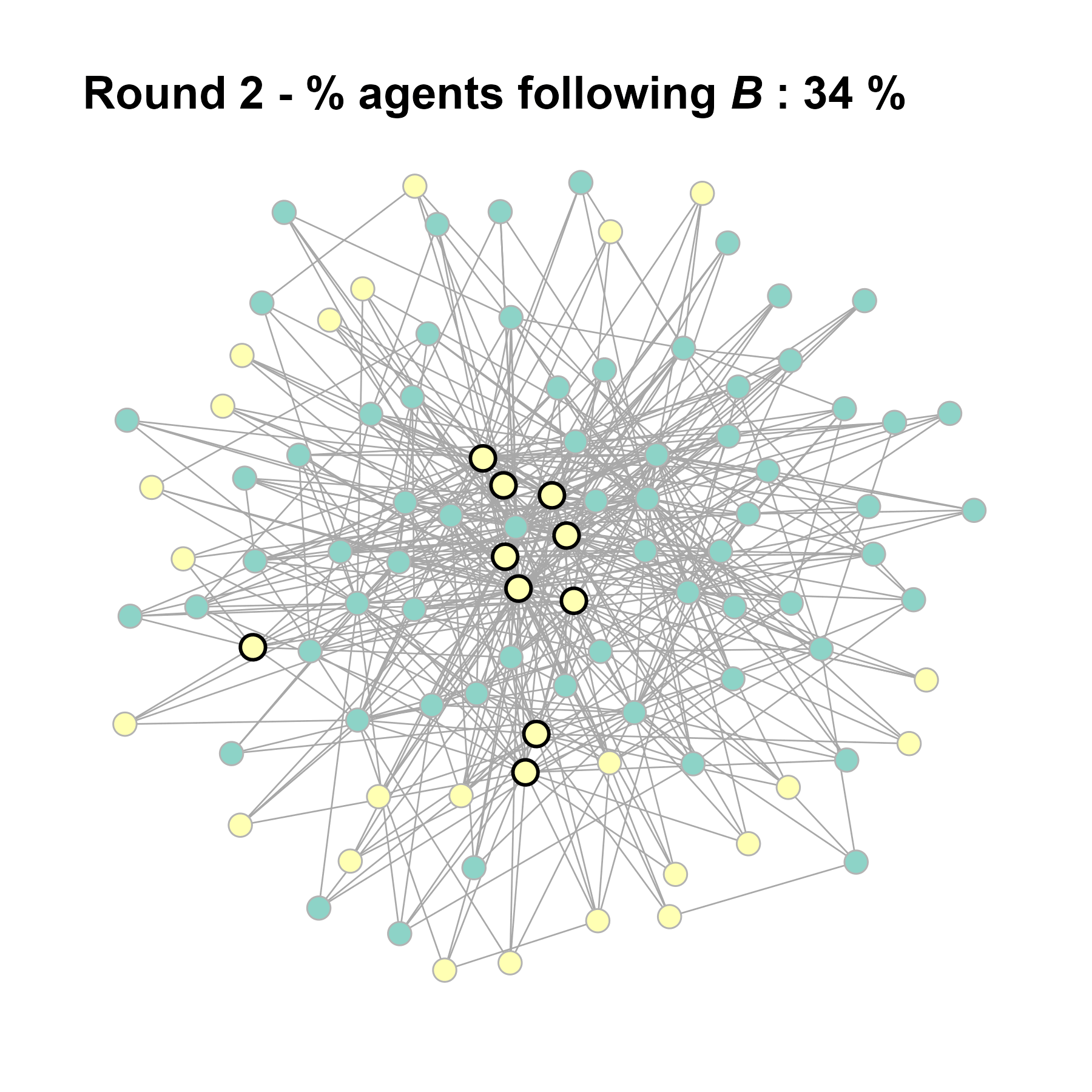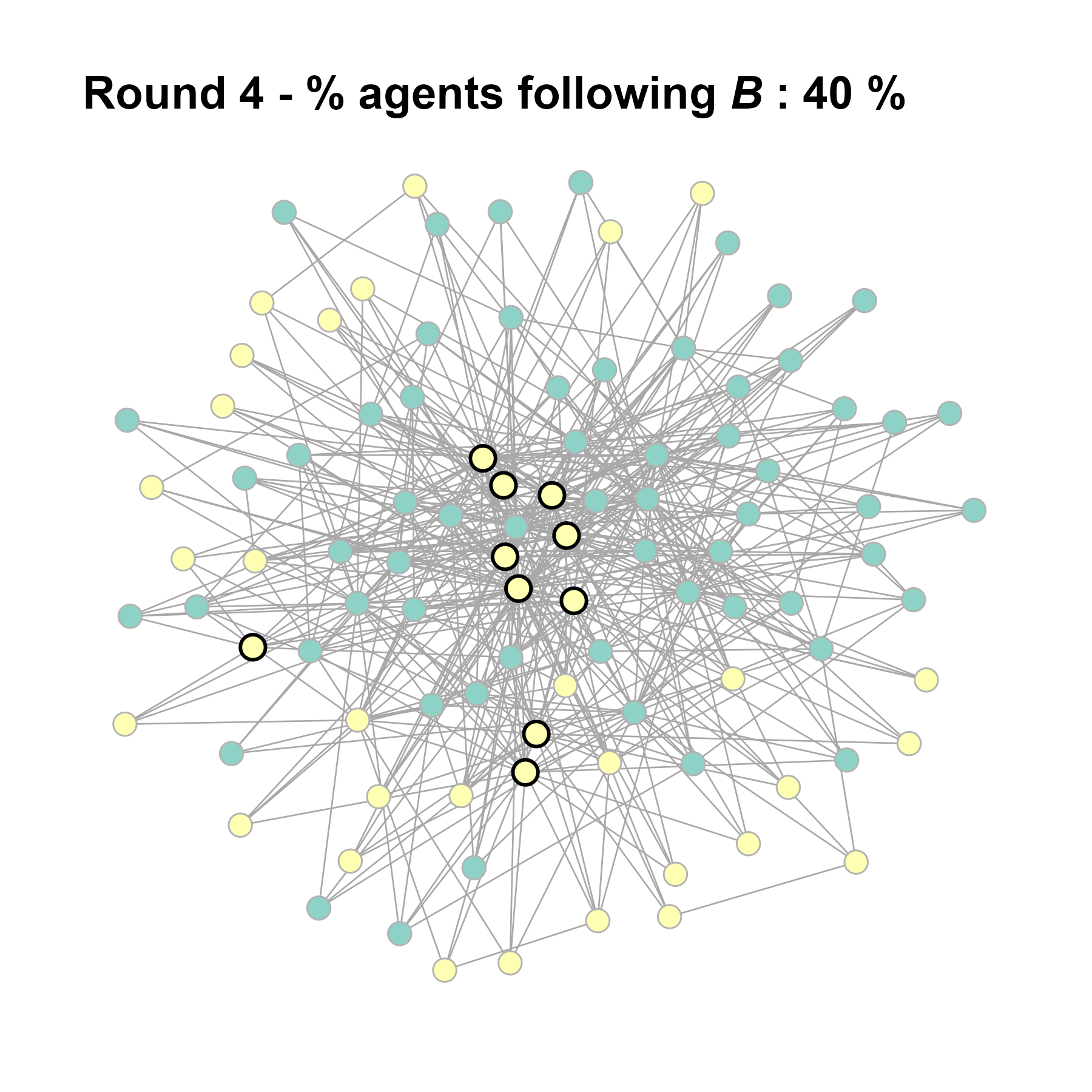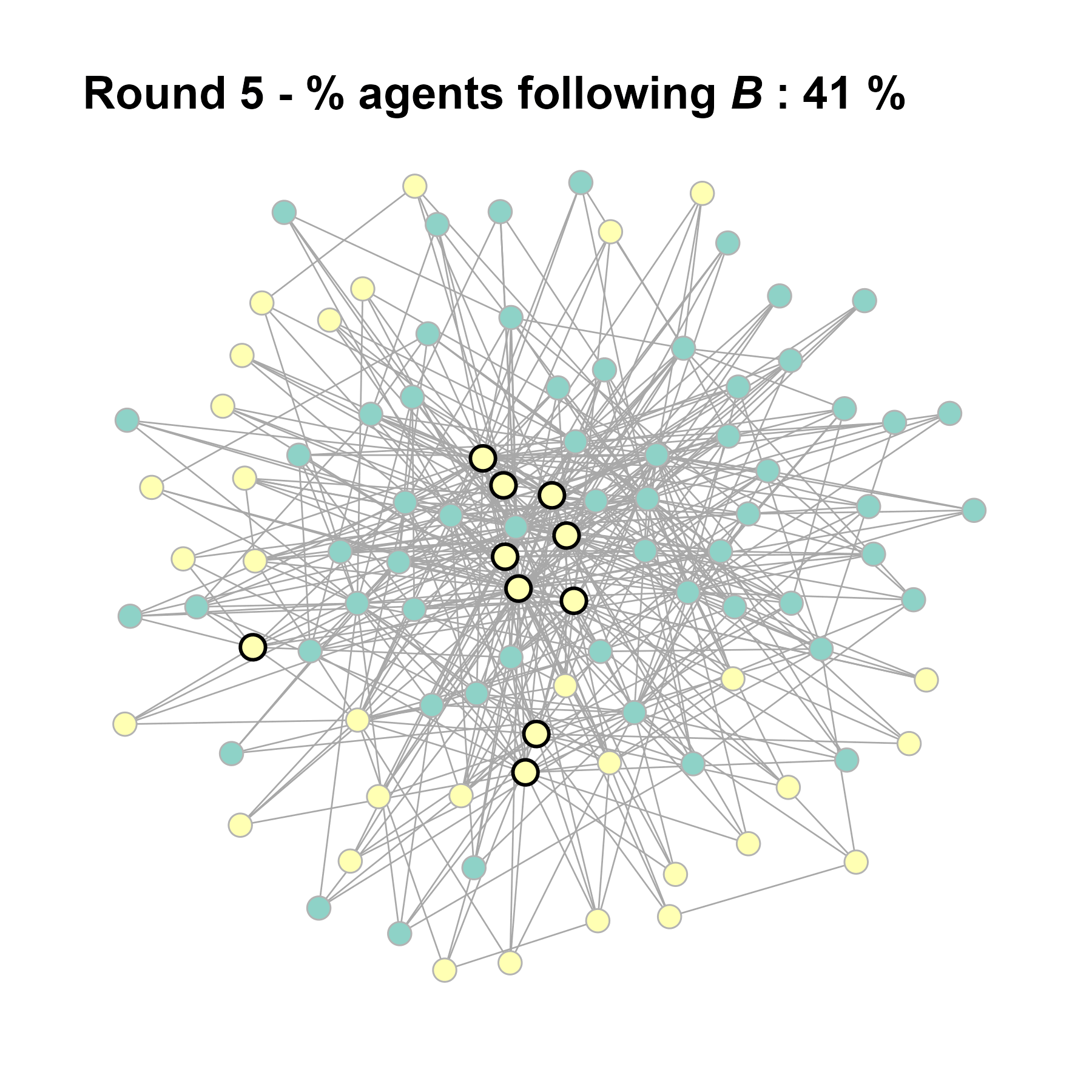
test <- run_one_seed(11, k_min = 4, k_max = 99, alpha = 2.1, rho = 0.5, r = -0.4)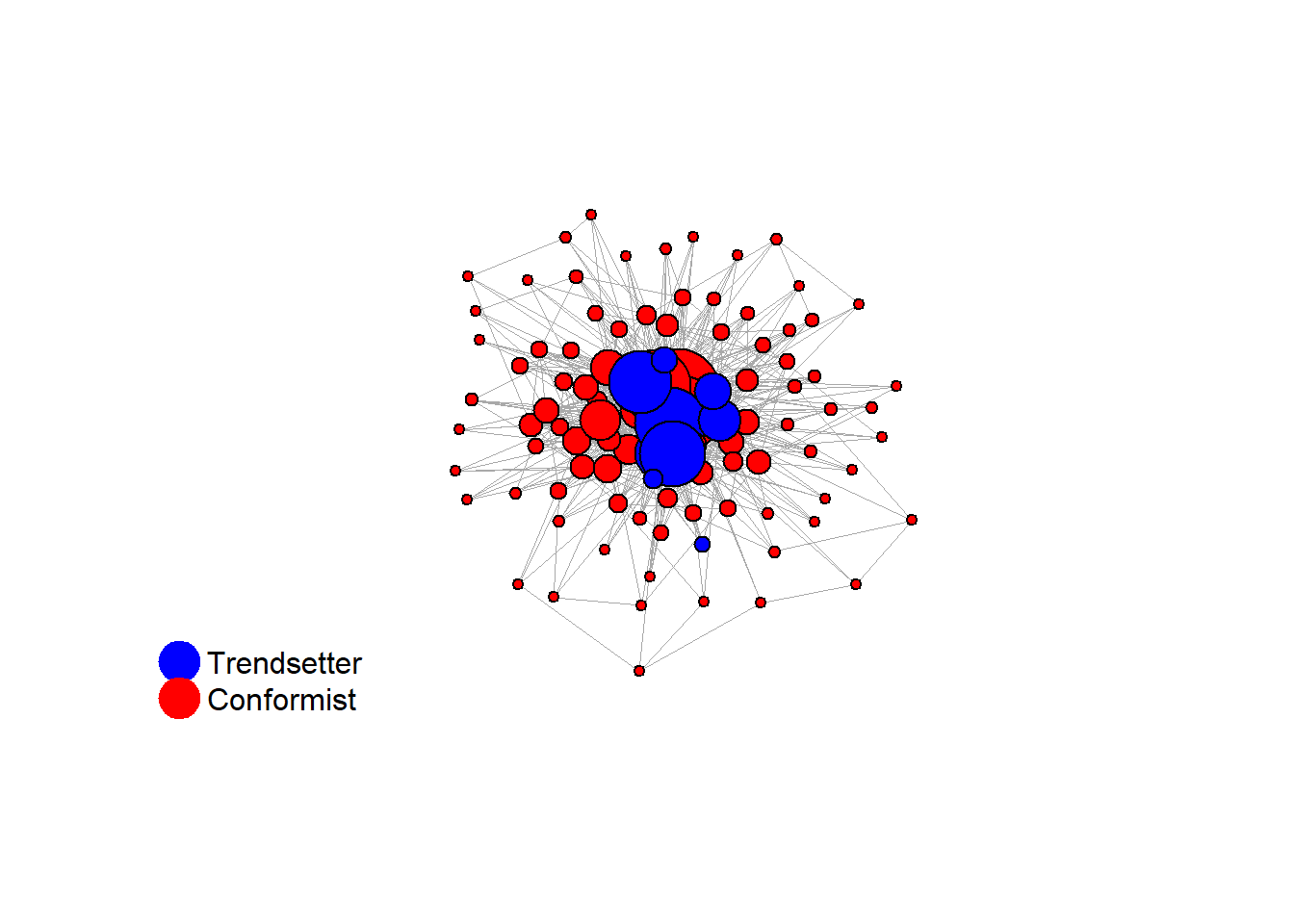
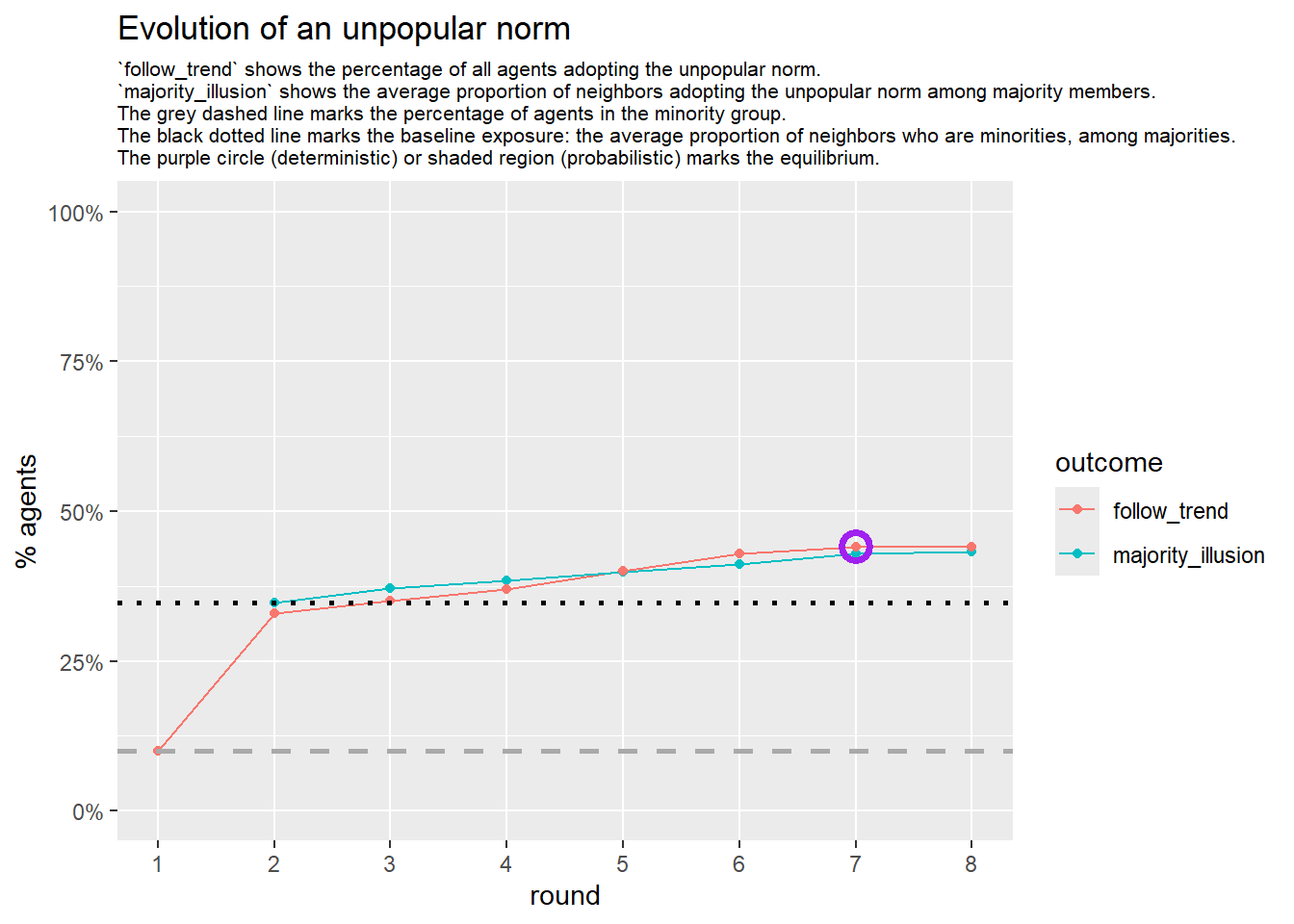
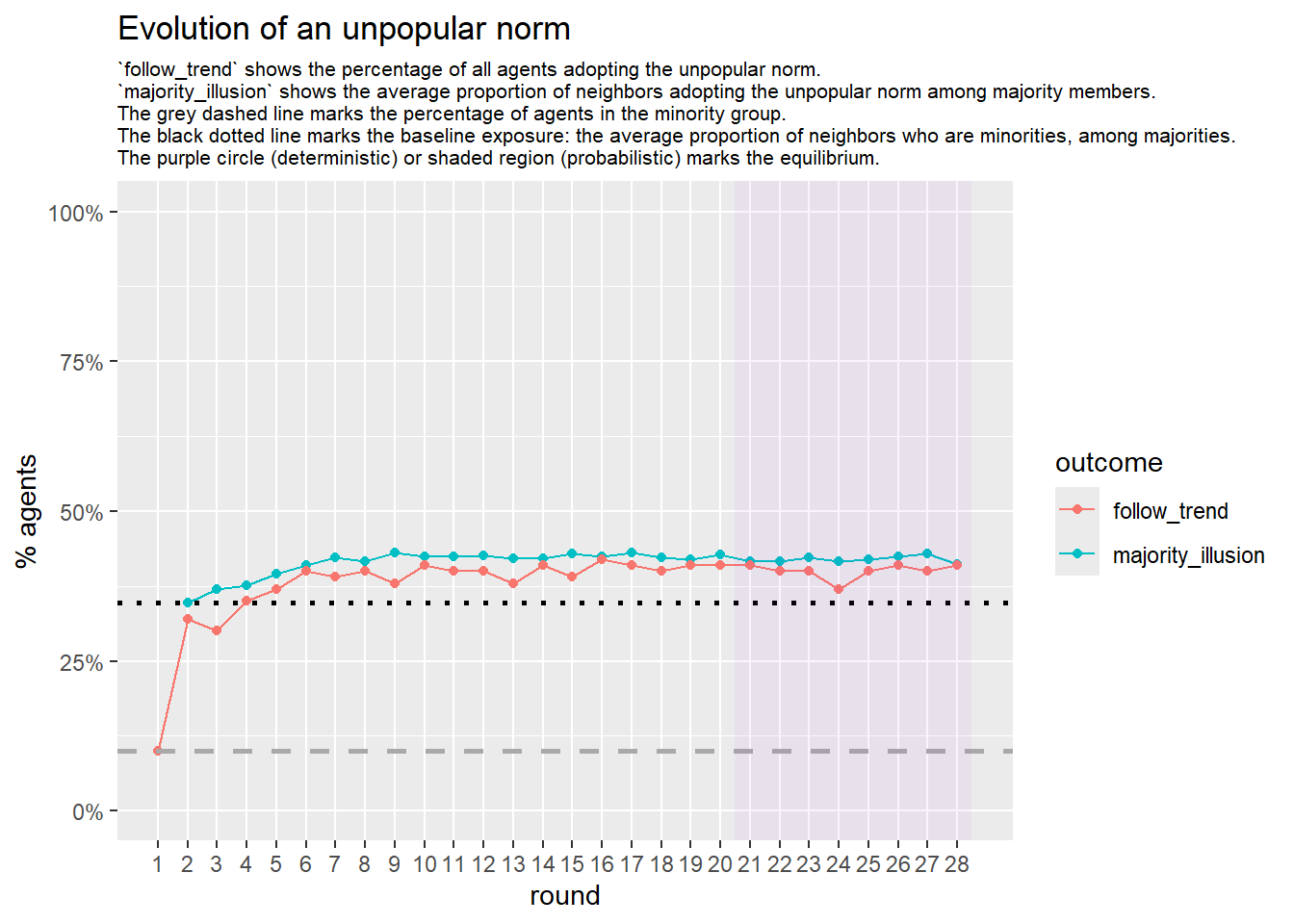
seed = seed + 1knitr::include_graphics(paste0("./figures/animation_network_", seed, ".gif"))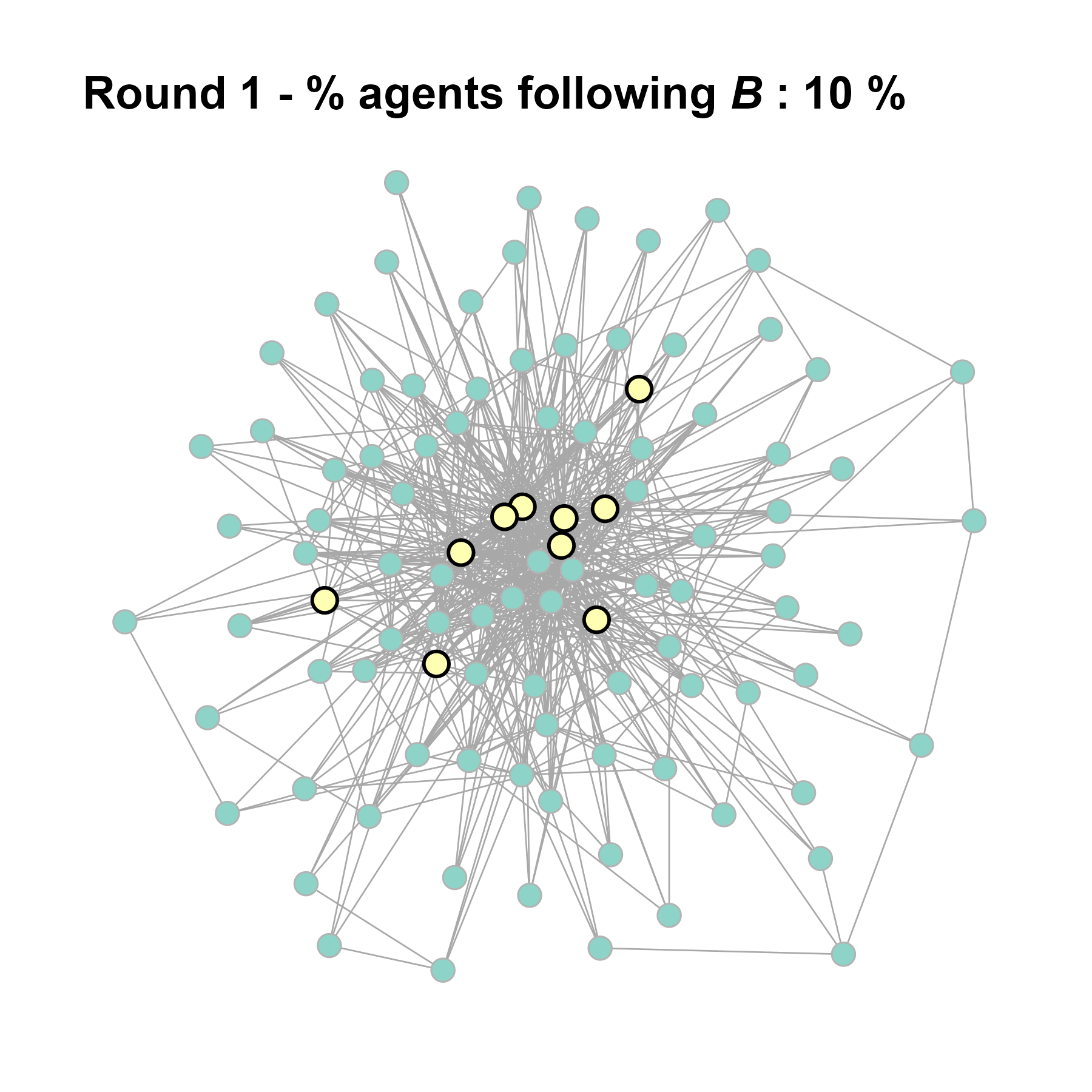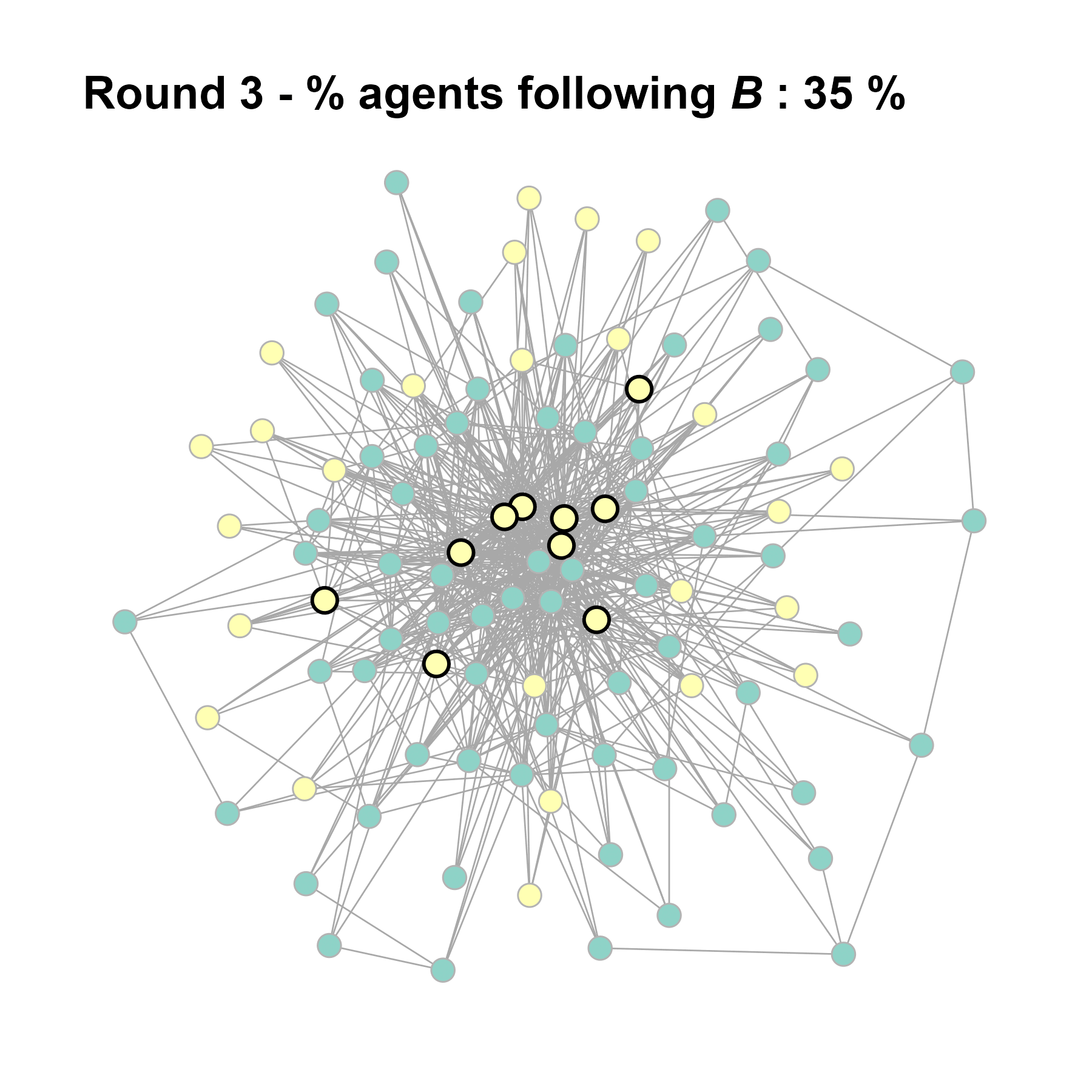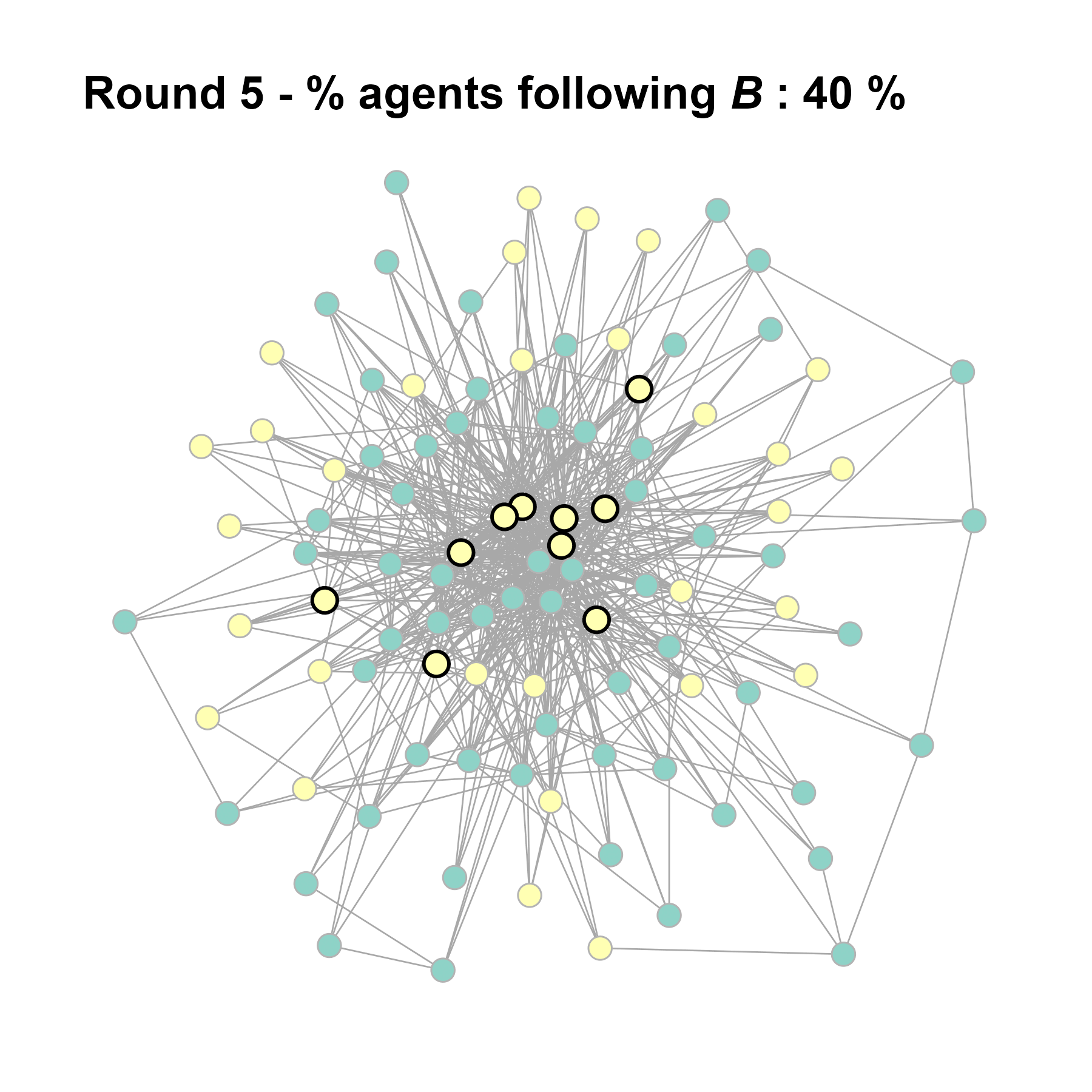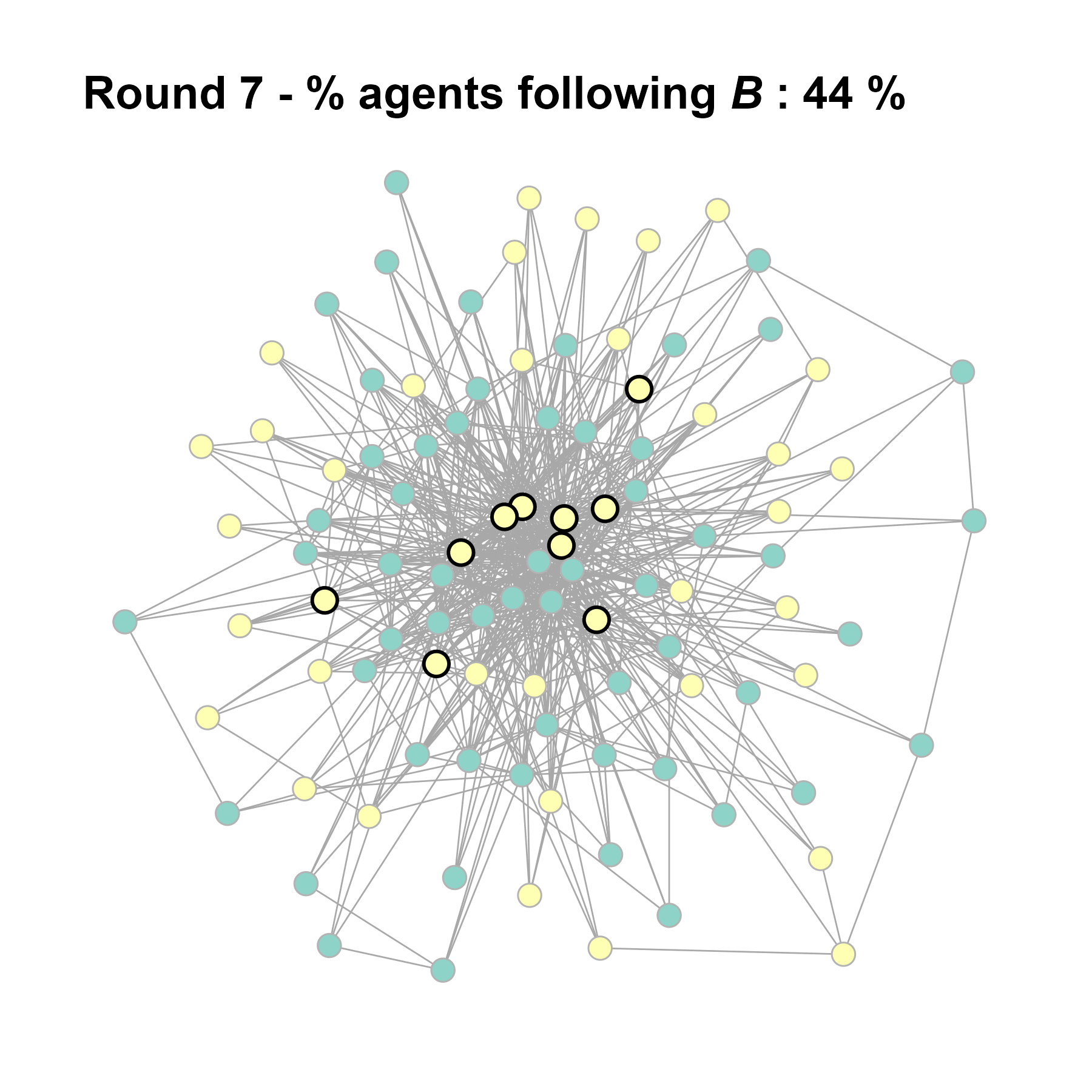
test <- run_one_seed(12, k_min = 4, k_max = 99, alpha = 2.1, rho = 0.5, r = -0.4)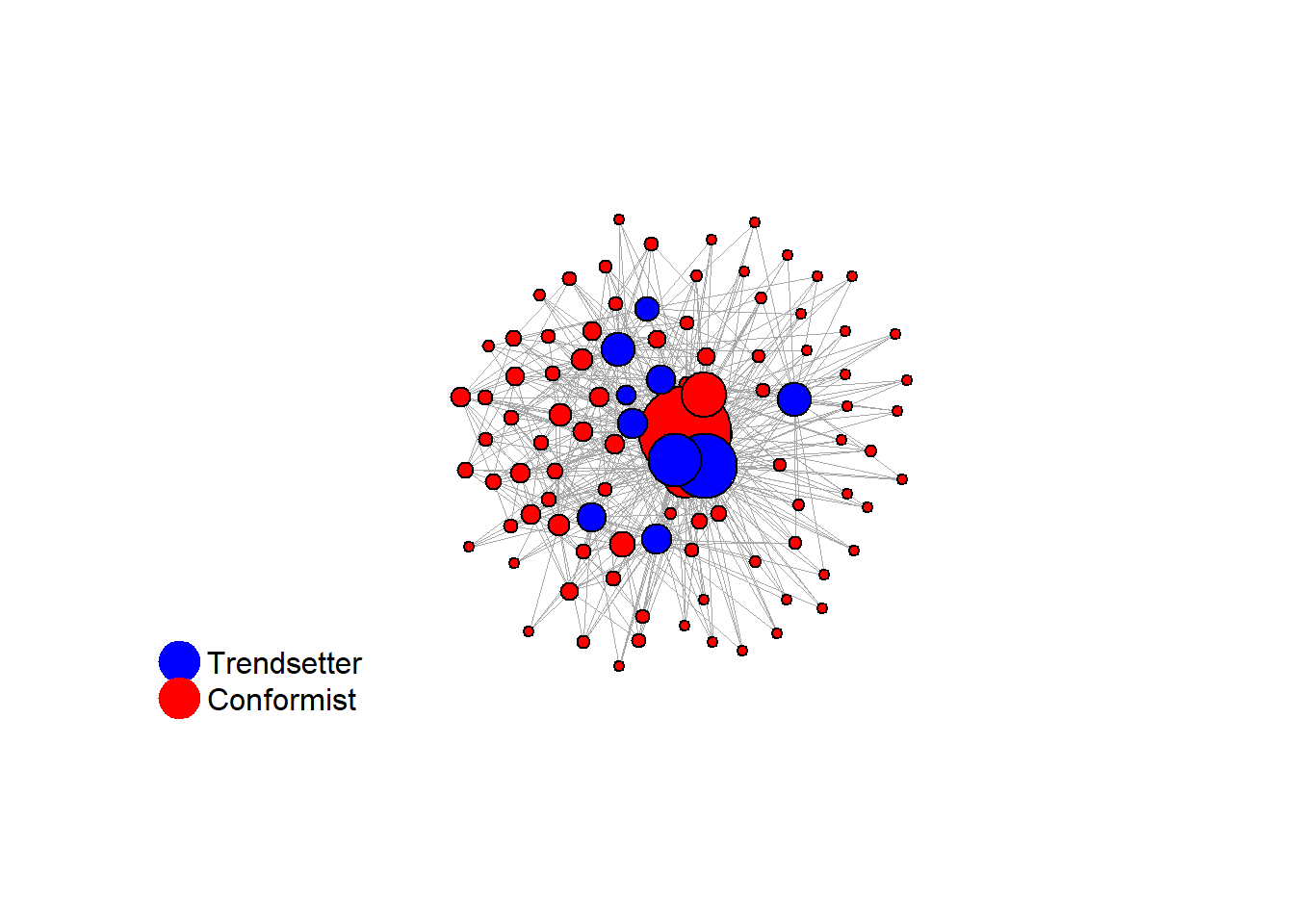
seed = seed + 1knitr::include_graphics(paste0("./figures/animation_network_", seed, ".gif"))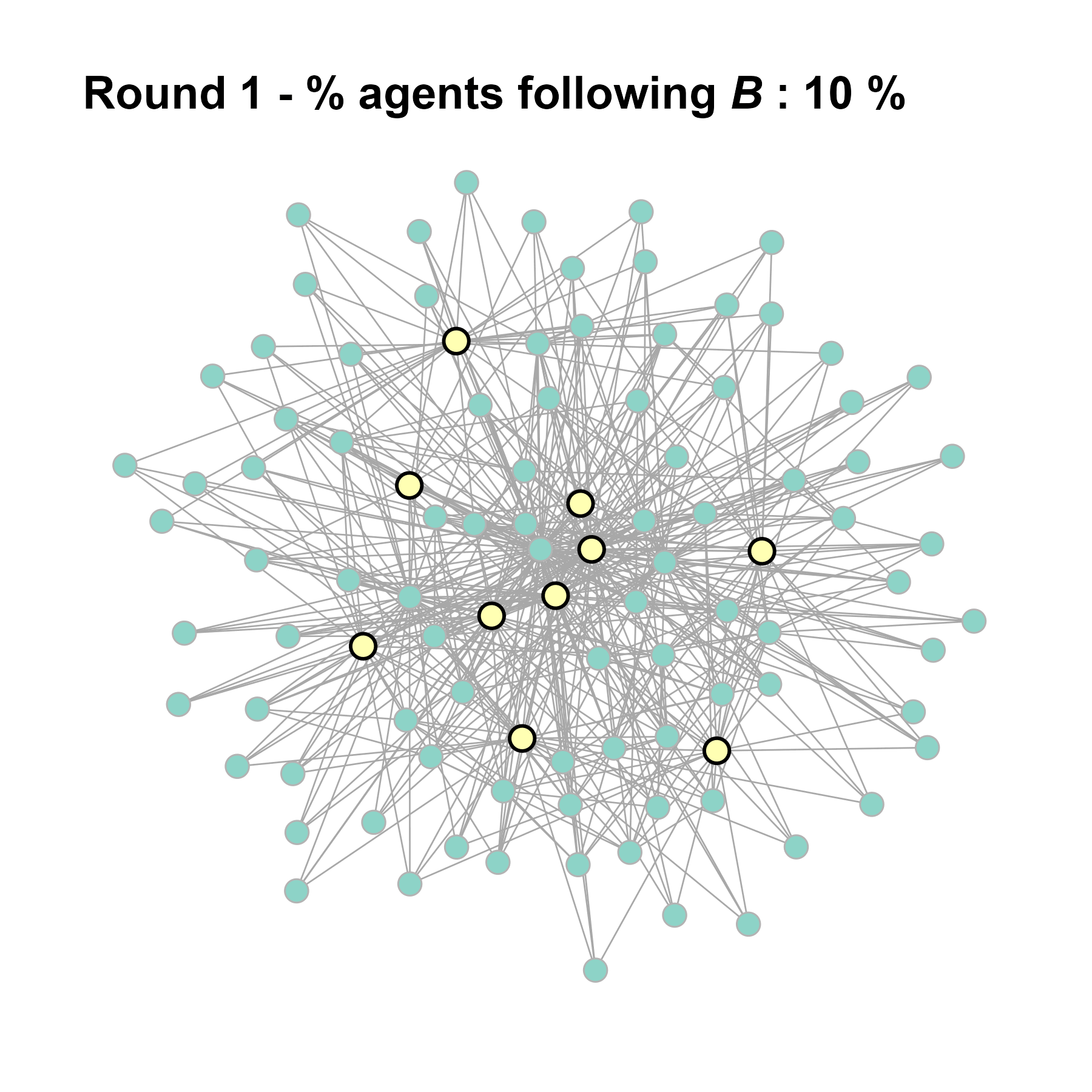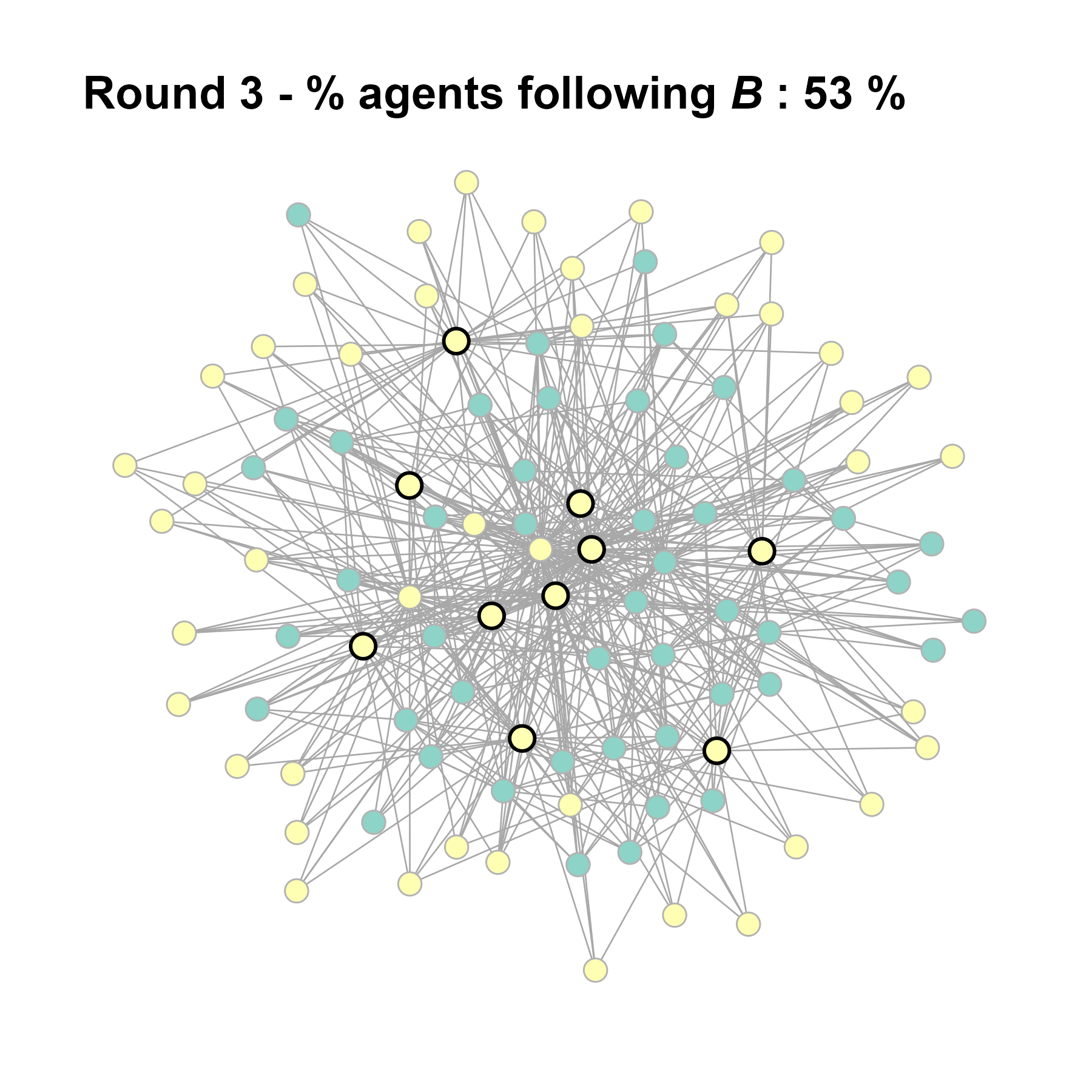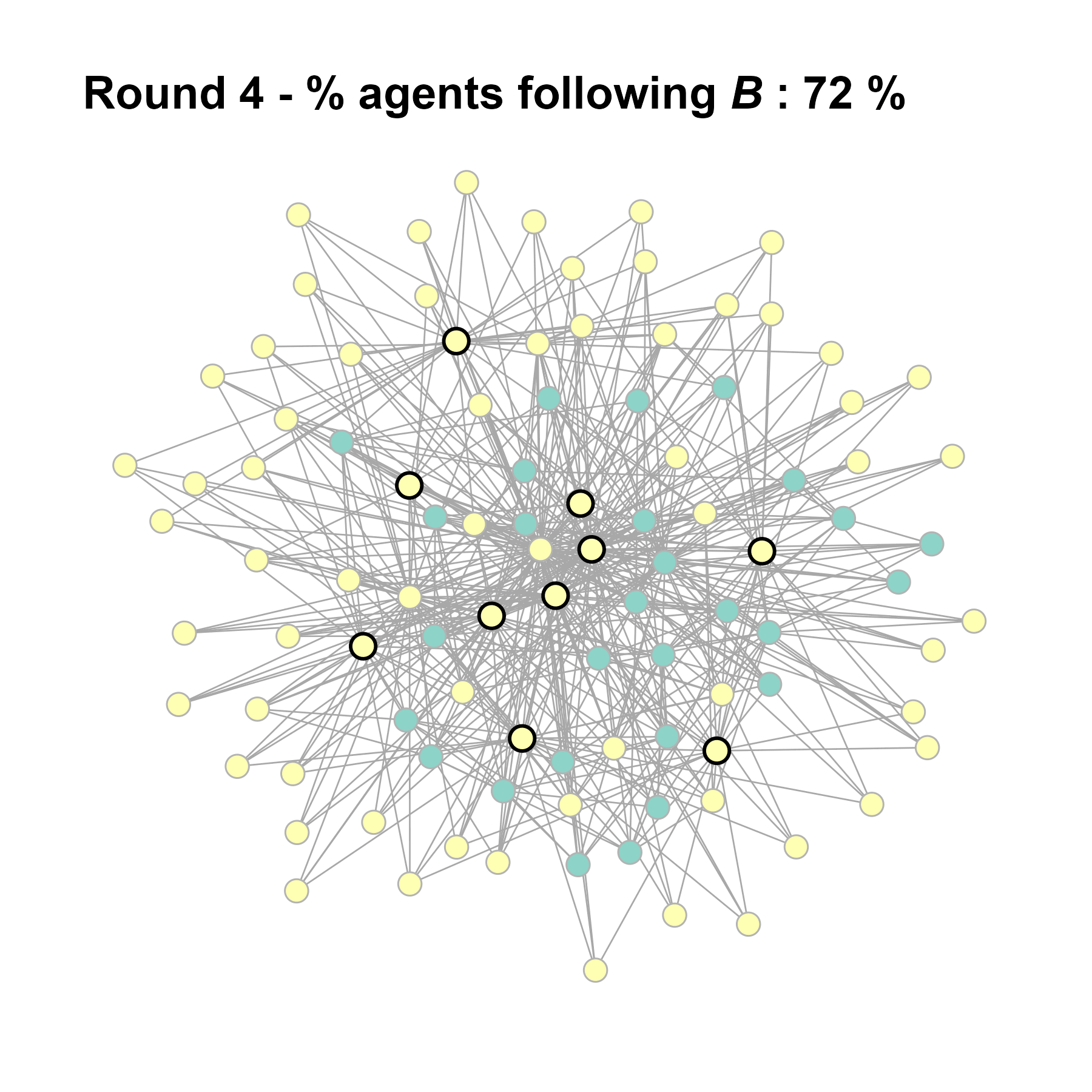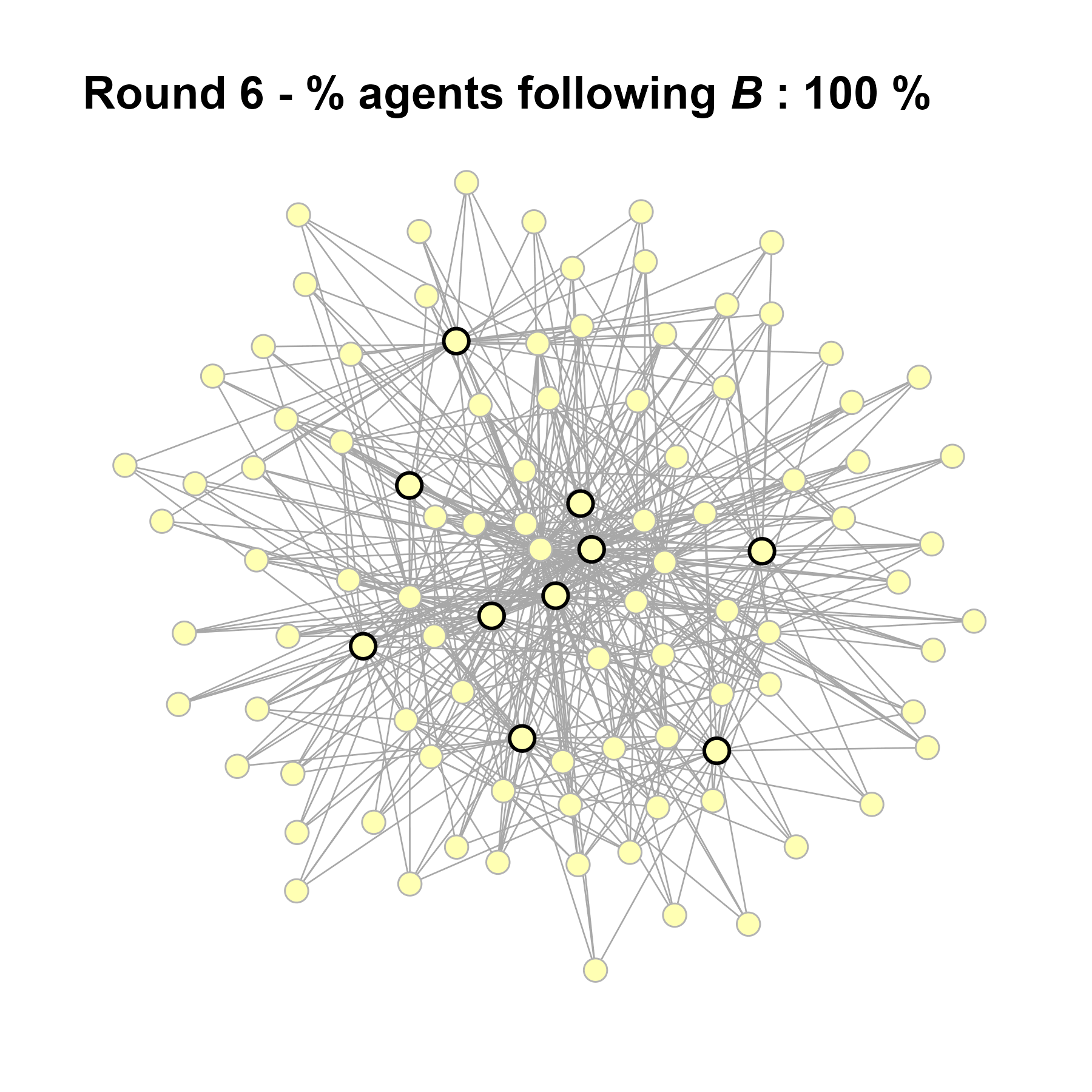
test <- run_one_seed(13, k_min = 4, k_max = 99, alpha = 2.1, rho = 0.5, r = -0.4)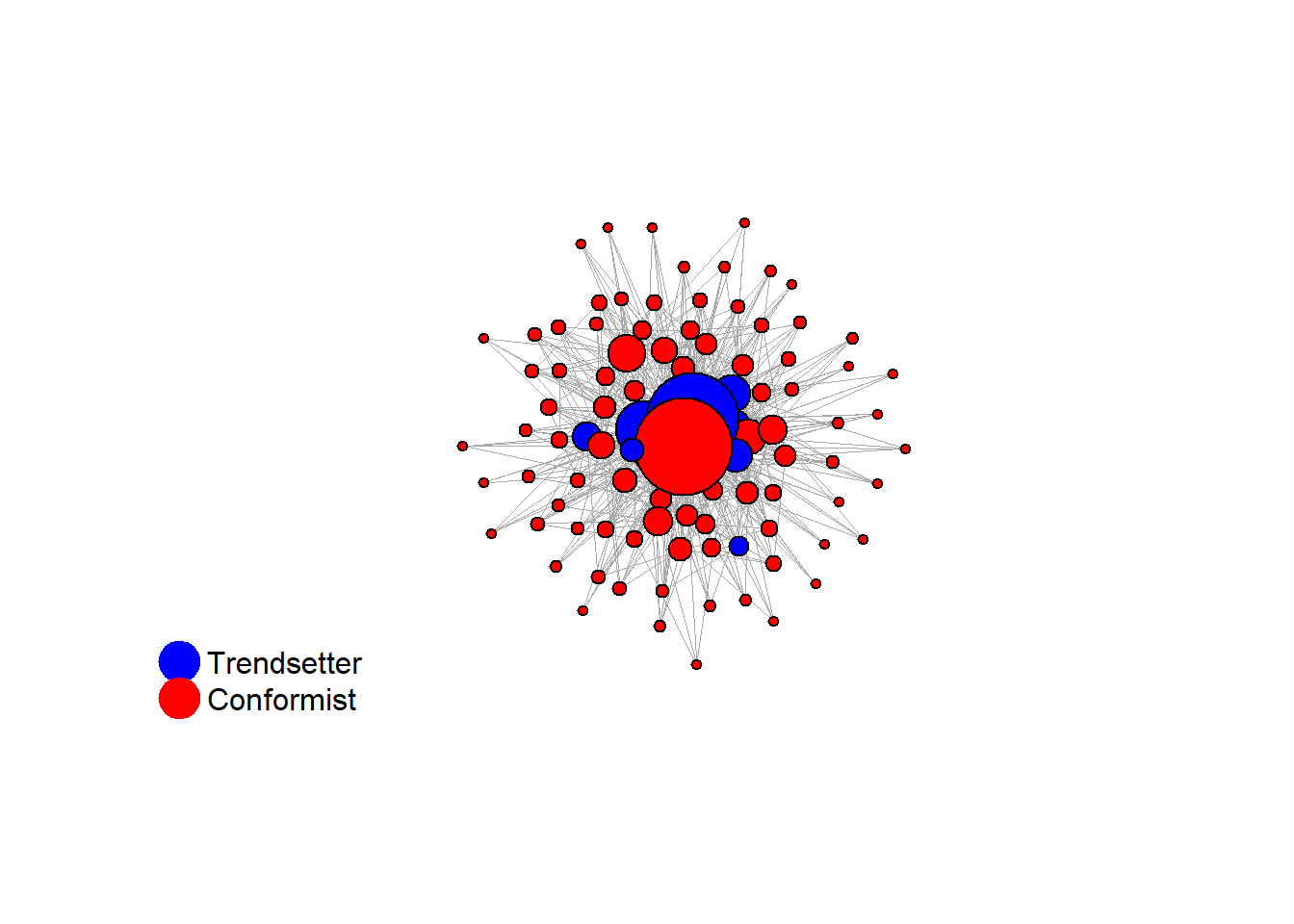
seed = seed + 1knitr::include_graphics(paste0("./figures/animation_network_", seed, ".gif"))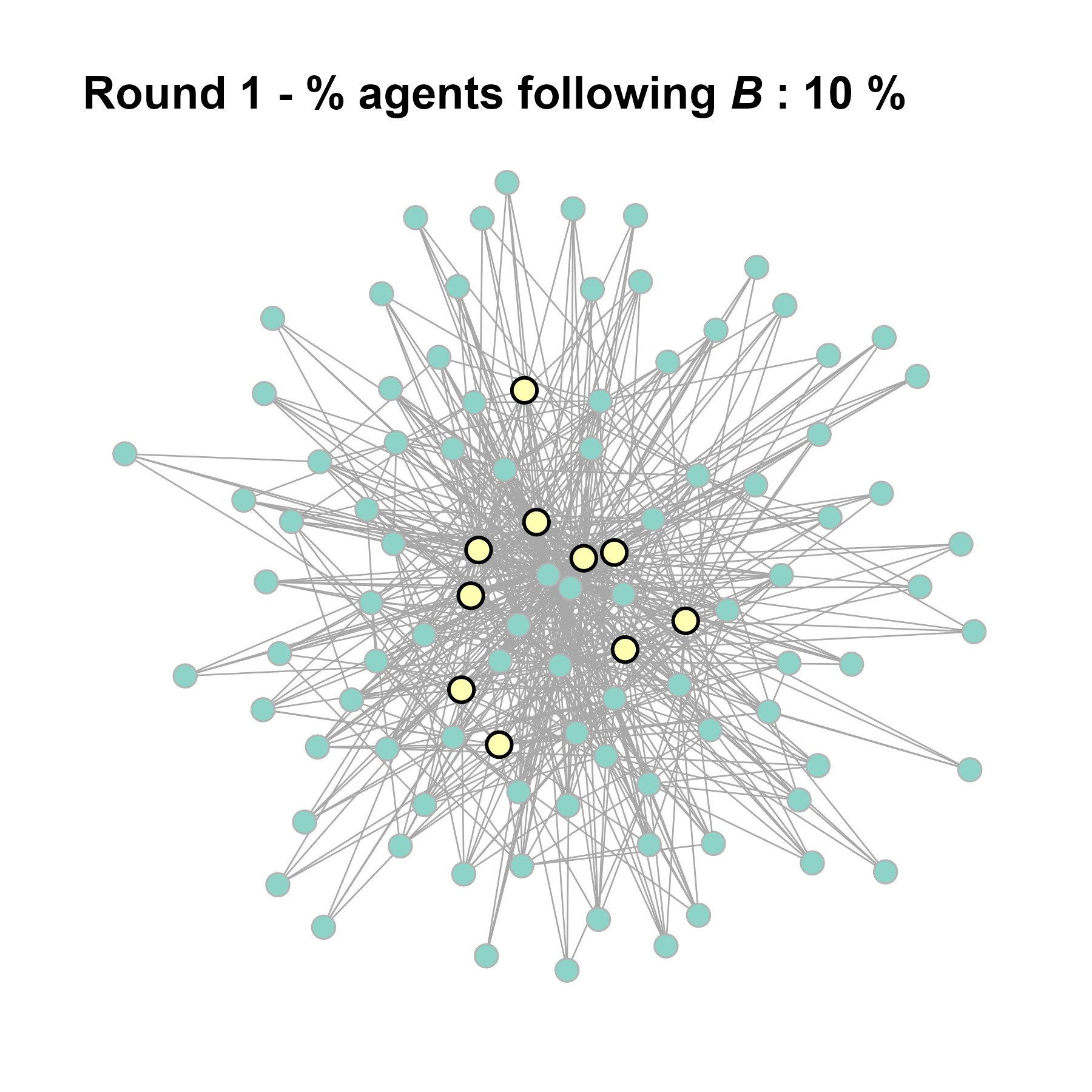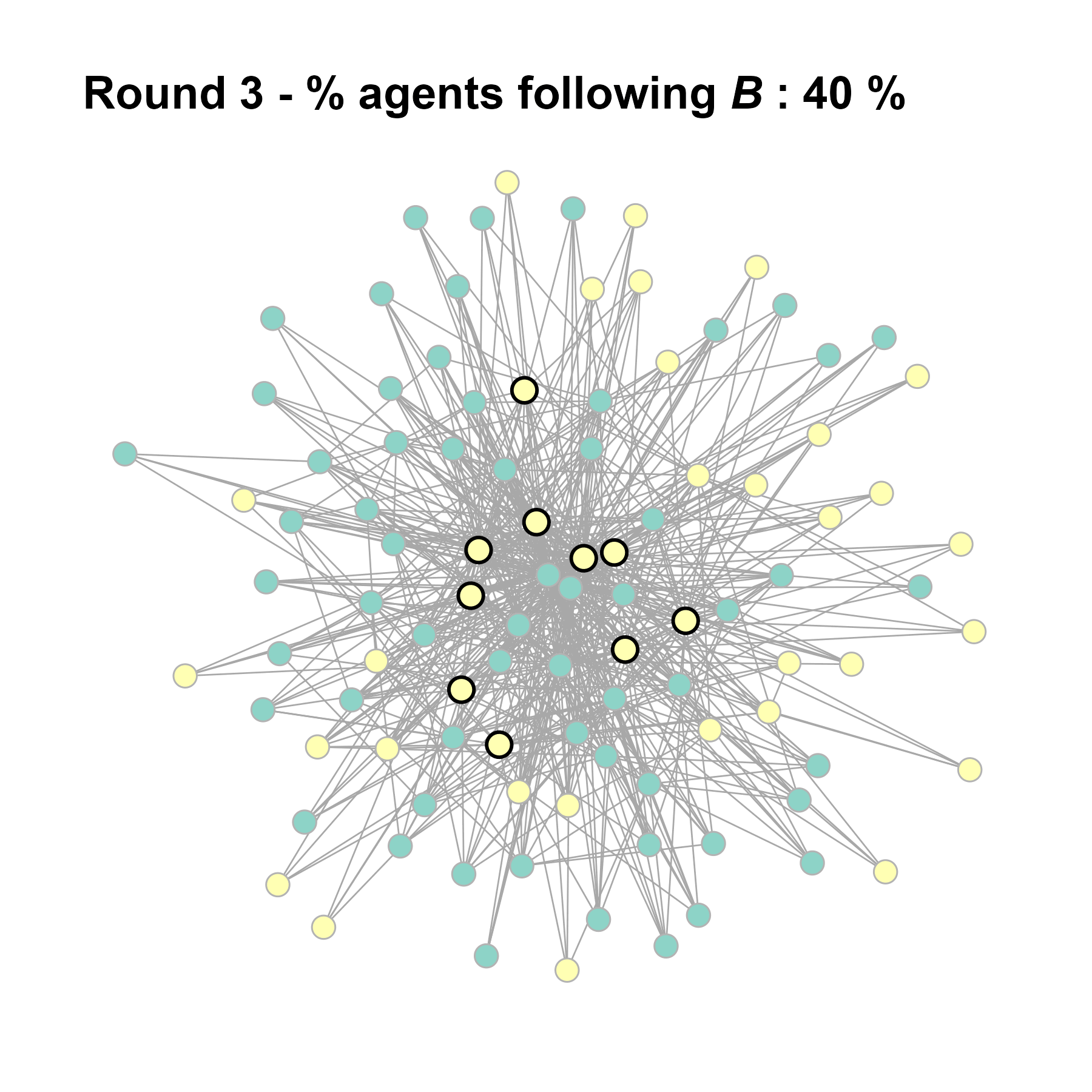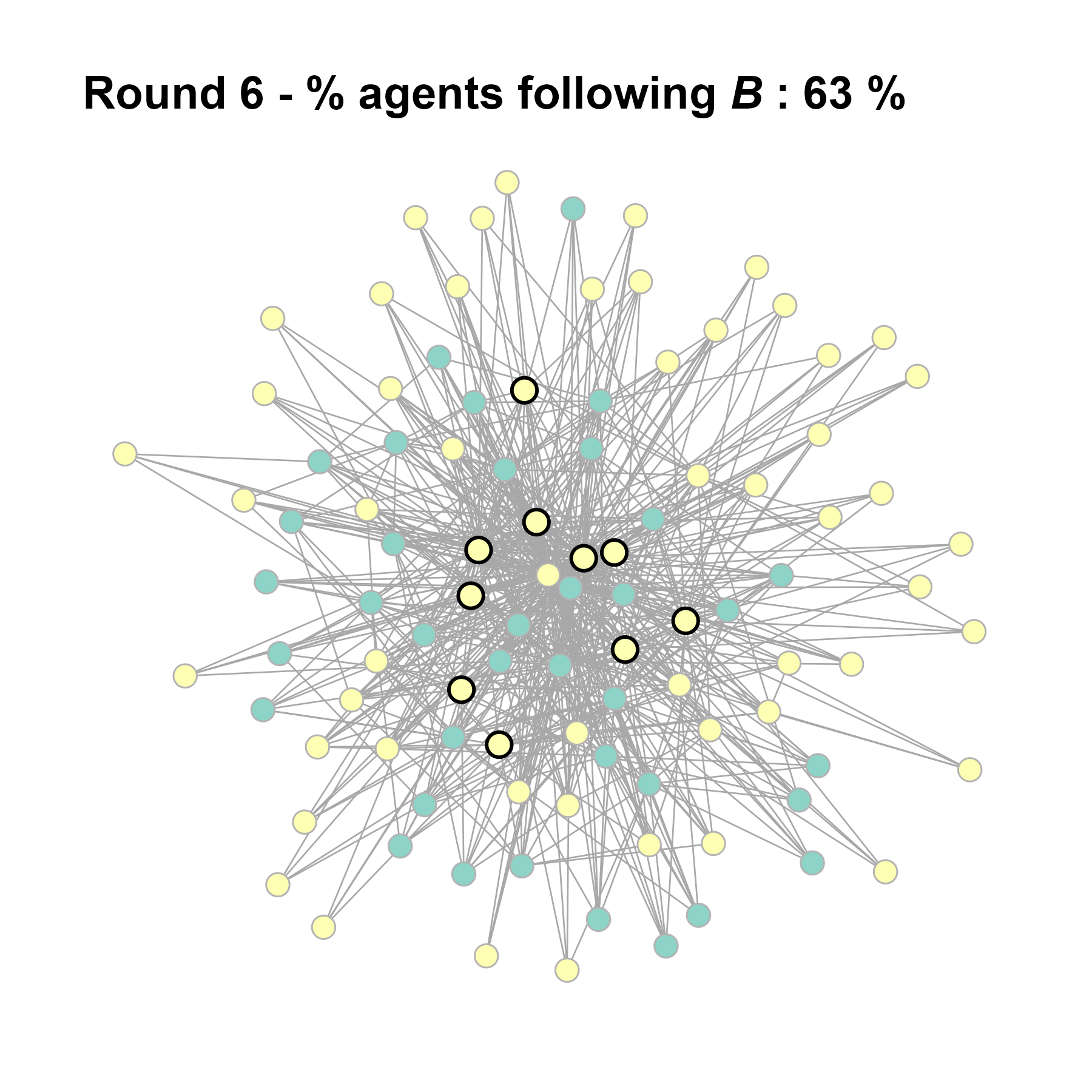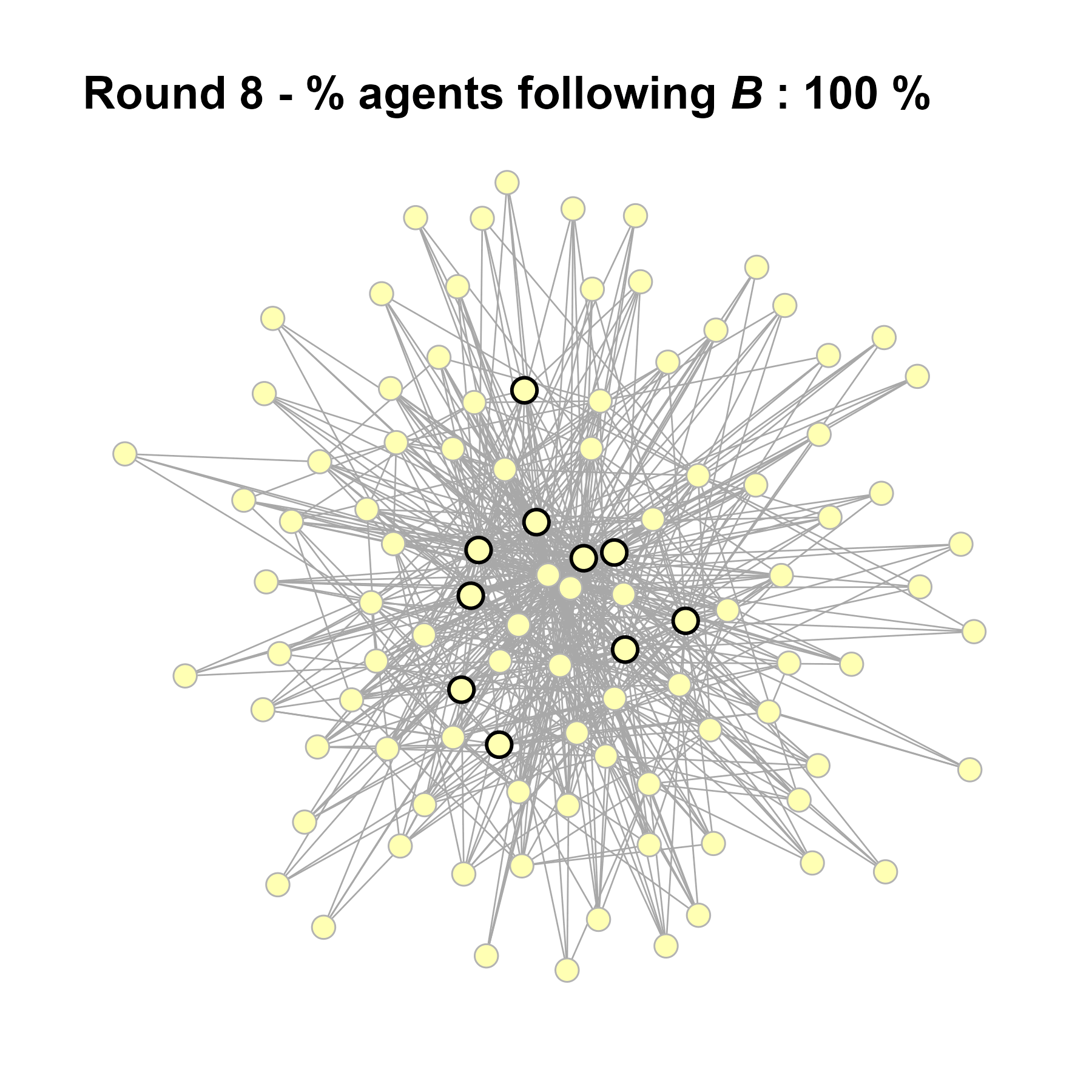
test <- run_one_seed(14, k_min = 4, k_max = 99, alpha = 2.1, rho = 0.5, r = -0.4)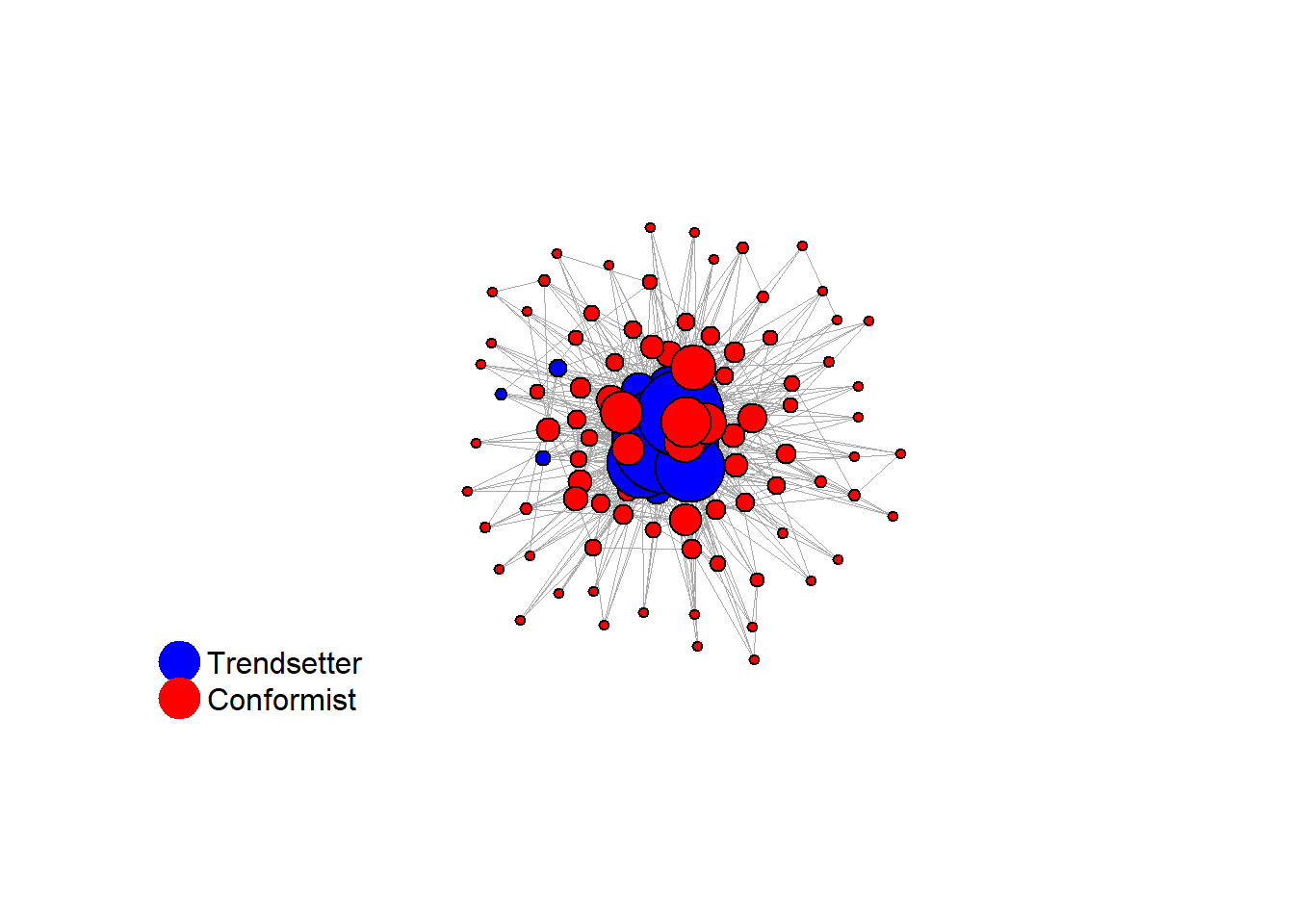
seed = seed + 1knitr::include_graphics(paste0("./figures/animation_network_", seed, ".gif"))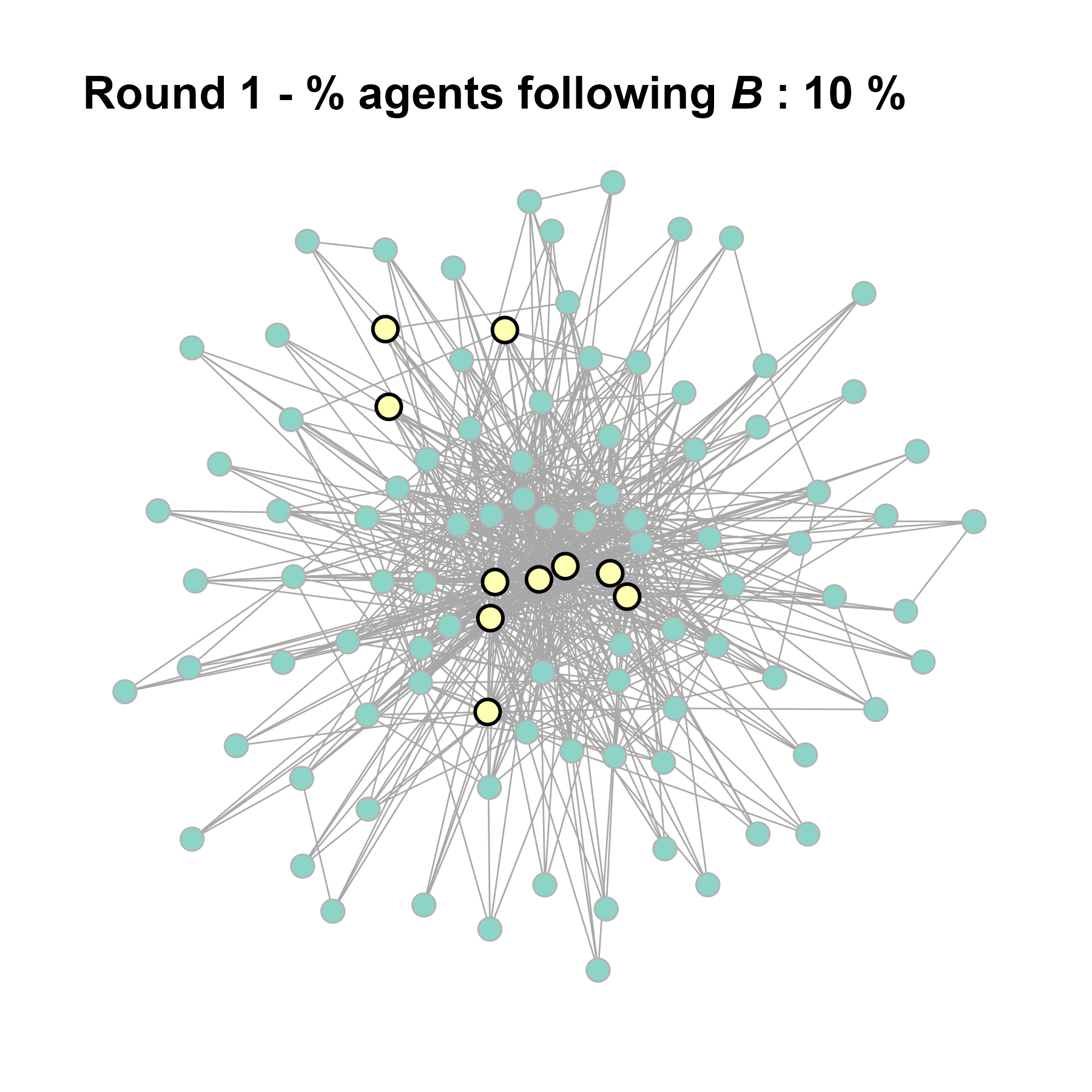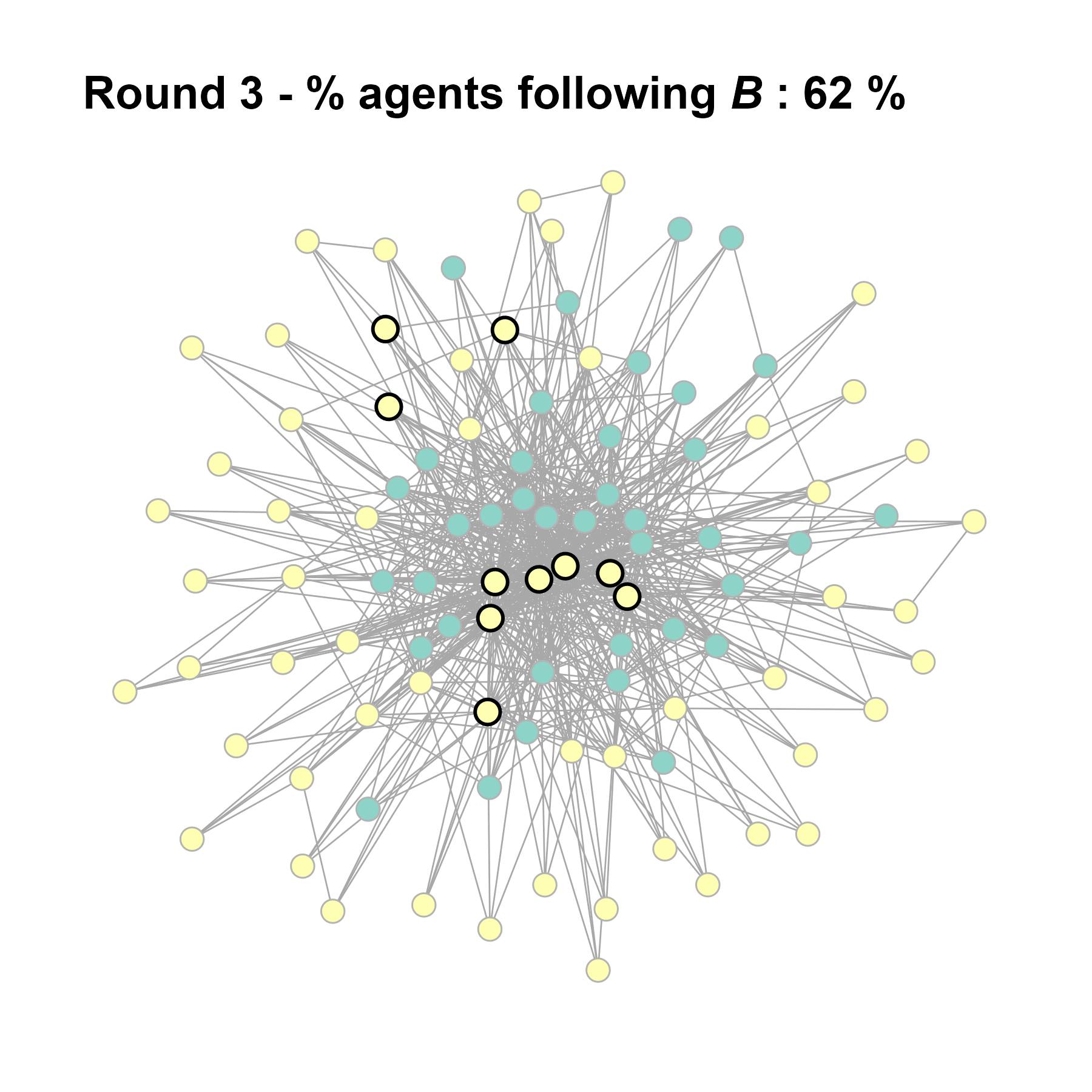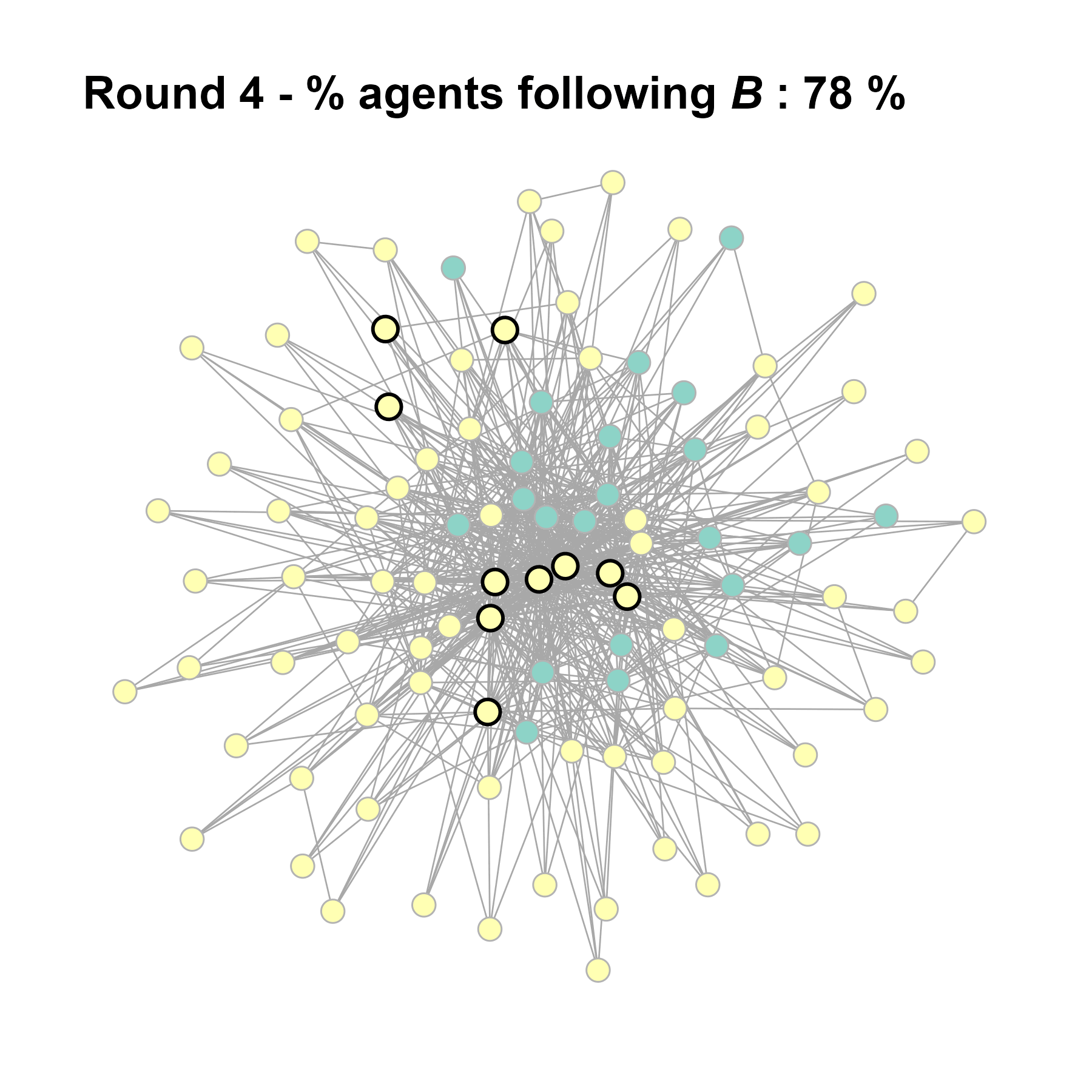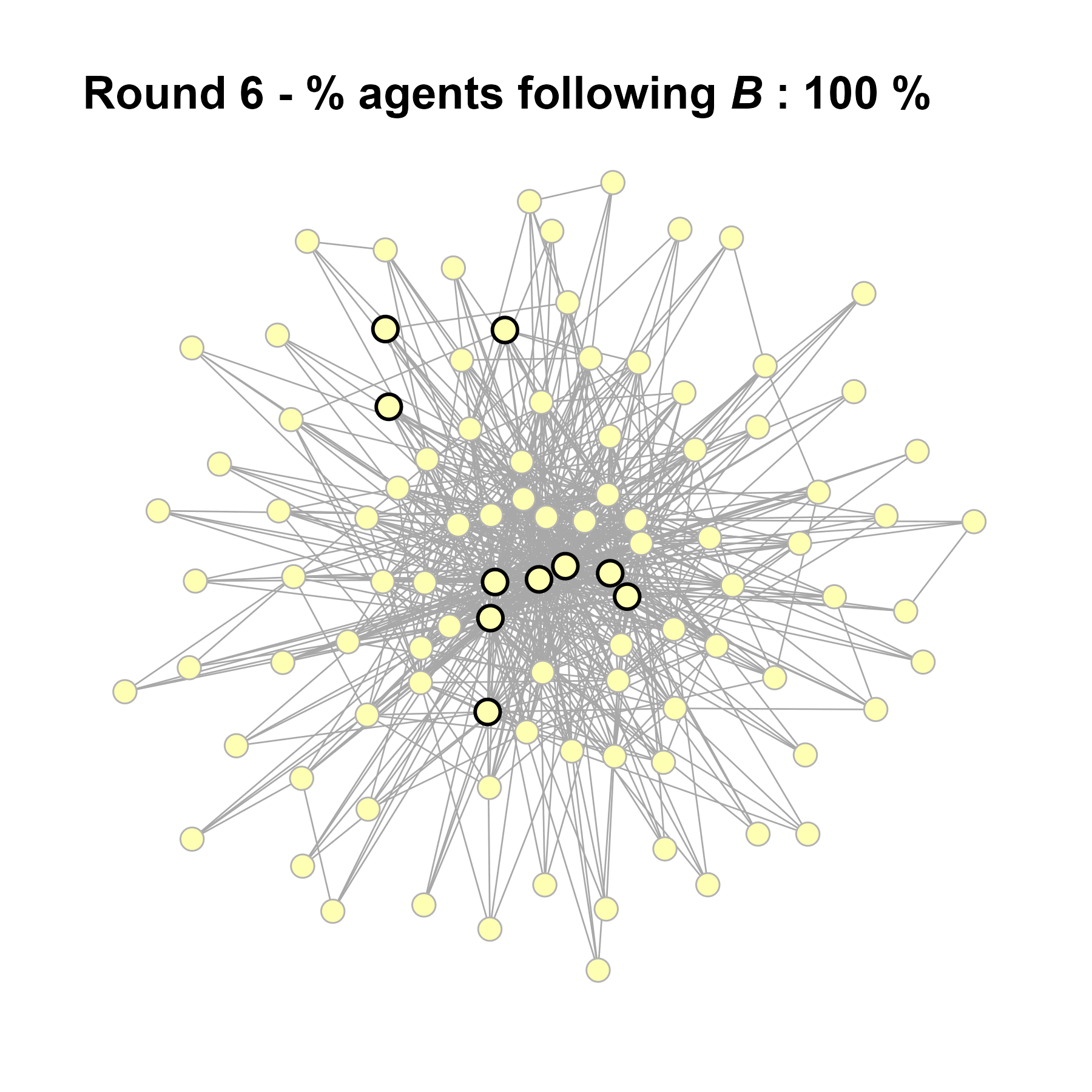
2.2 2. small-worlds
run_one_sw <- function(
i,
base_seed = 1253281,
model = "watts-strogatz",
beta = 0,
nei = 3,
clique_size = 5,
pmin = 0.1,
params = list(s = 15, e = 10, w = 40, z = 50, lambda1 = 5, lambda2 = 1.8),
# retrieve network
return_network = FALSE
) {
# derived seed for this run
seed_i <- base_seed + i
set.seed(seed_i)
if (model == "watts-strogatz") {
network <- sample_smallworld(dim = 1, size = 100, nei = 3, p = beta) }
else if (model == "caveman") {
network <- simulate_caveman(n = 100, clique_size = clique_size)
}
V(network)$role <- sample(
c(rep("trendsetter", round(100*pmin)), rep("conformist", 100-round(100*pmin)))
)
final_network <- network
# --- stats ---
stats <- list(
run = i,
seed = seed_i,
num_nodes = vcount(final_network),
num_edges = ecount(final_network),
avg_degree = mean(degree(final_network)),
sd_degree = sd(degree(final_network)),
net_density = edge_density(final_network),
net_diameter = diameter(final_network, directed = FALSE, unconnected = TRUE),
avg_path_len = average.path.length(final_network, directed = FALSE),
clust_coeff = transitivity(final_network, type = "global"),
assort_deg = assortativity_degree(final_network),
deg_trait_cor = fdegtraitcor(final_network)$cor,
components = components(final_network)$no
)
stats_df <- data.frame(
Metric = names(stats),
Value = unlist(stats),
row.names = NULL
)
#print(stats_df)
fplot_graph(final_network, layout = layout.kamada.kawai(final_network))
# --- initial actions ---
V(final_network)$action <- ifelse(V(final_network)$role == "trendsetter", 1, 0)
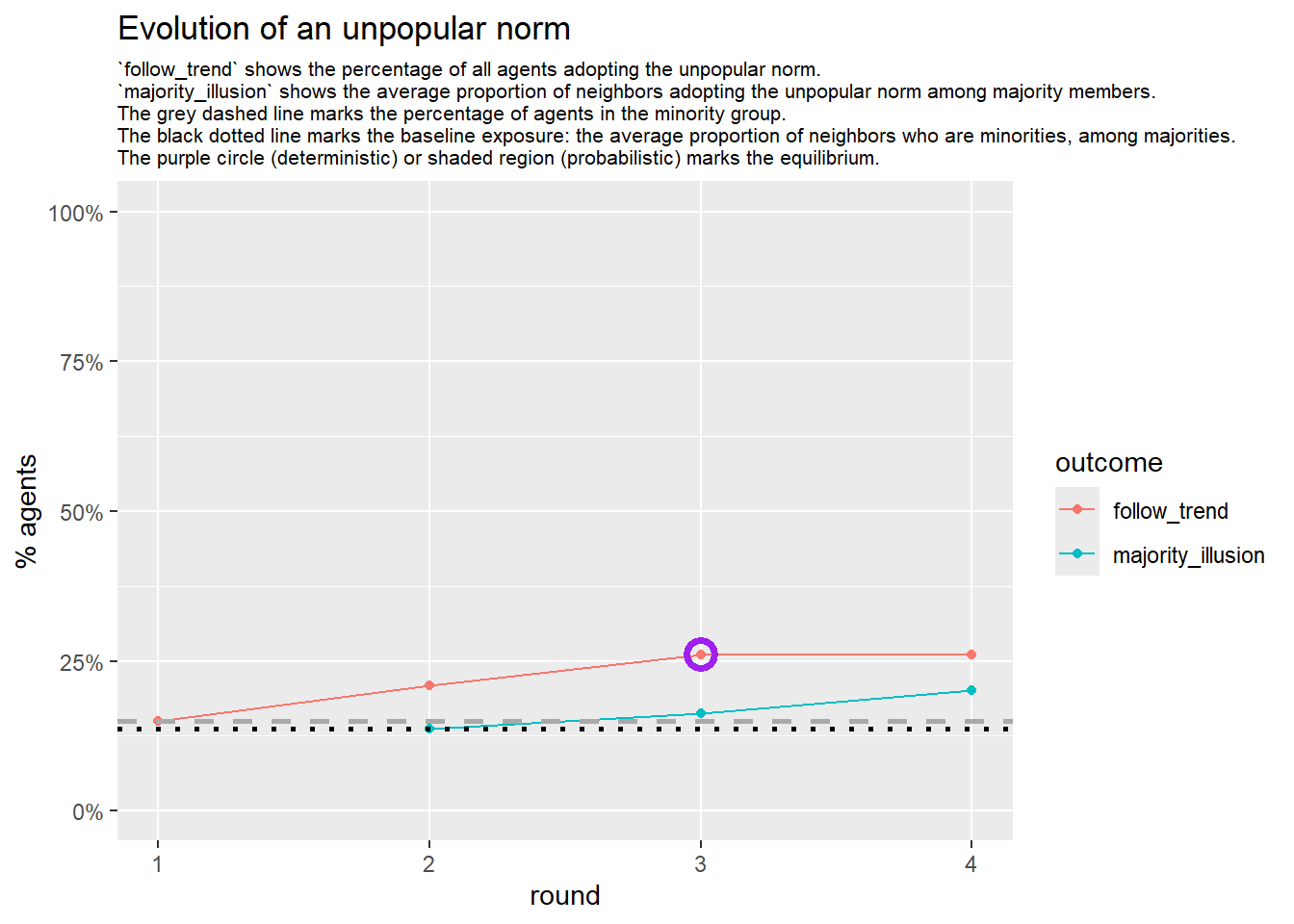
# --- deterministic simulation ---
sim_det <- fabm(
network = final_network,
params = params,
max_rounds = 50,
mi_threshold = 0.49,
choice_rule = "deterministic",
plot = TRUE,
histories = TRUE
)
# generate the gif for the current network
gif_filename <- paste0("./figures/animation_network_", seed_i, ".gif")
gif_path <- fnetworkgif(final_network, sim_det$decision_history, rounds = sim_det$equilibrium$round, output_dir = "./figures")
# rename the gif to match the naming pattern
file.rename(gif_path, gif_filename)
if (!is.null(sim_det$plot)) {
print(sim_det$plot)
}
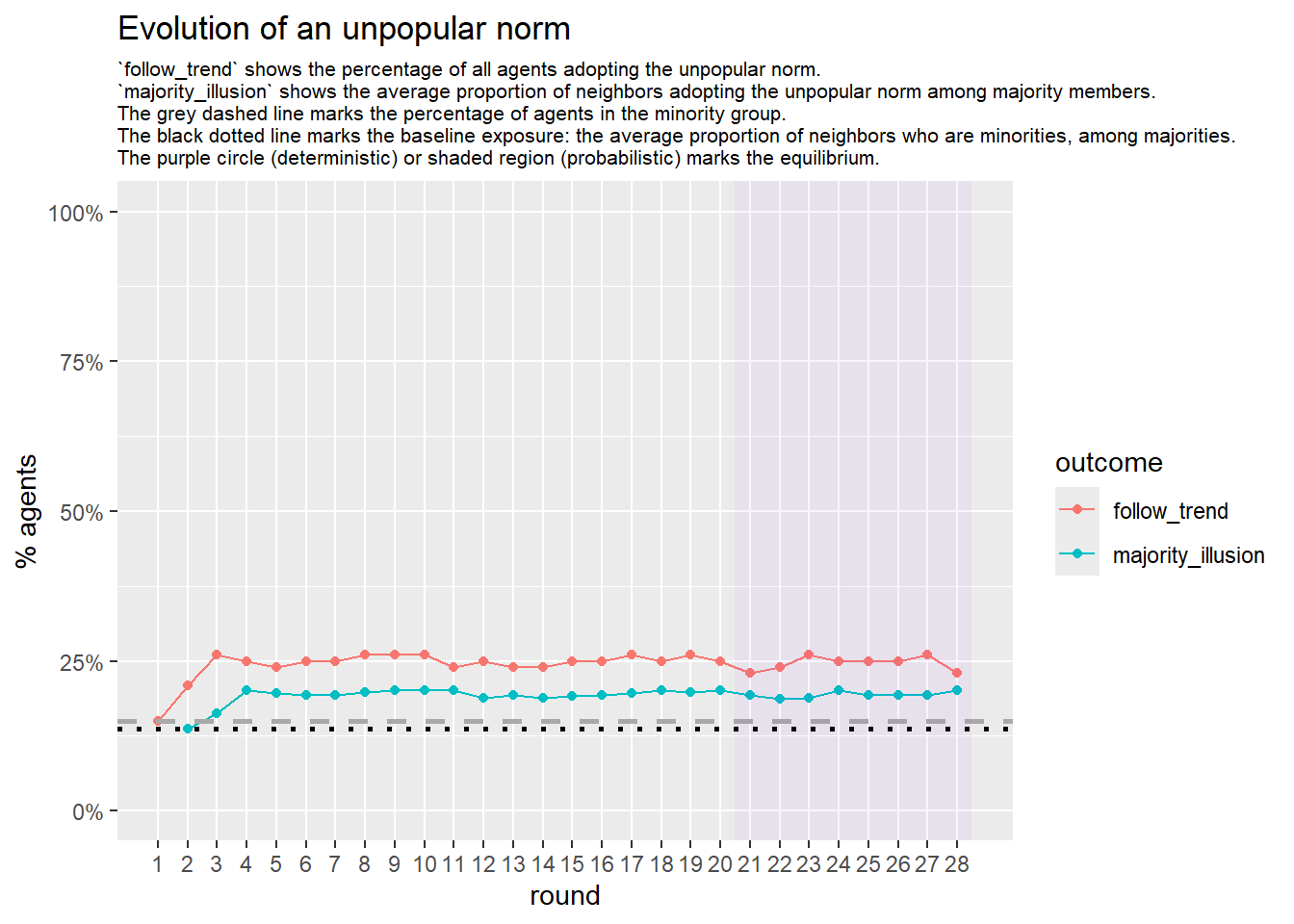
# --- probabilistic simulation ---
sim_prob <- fabm(
network = final_network,
params = params,
max_rounds = 100,
mi_threshold = 0.49,
choice_rule = "probabilistic",
stable_window = 8, # the length of the window of adoption values
required_stable_rounds = 20, # number of windows needed to declare equilibrium
plot = TRUE
)
if (!is.null(sim_prob$plot)) {
print(sim_prob$plot)
}
result <- list(
segregation_det = sim_det$equilibrium$segregation,
segregation_prob = sim_prob$equilibrium$segregation,
stats = stats
)
if (return_network) {
result$network <- final_network
}
result
}# random
test <- run_one_sw(i = 15, beta = 1, pmin = 0.1, return_network = TRUE)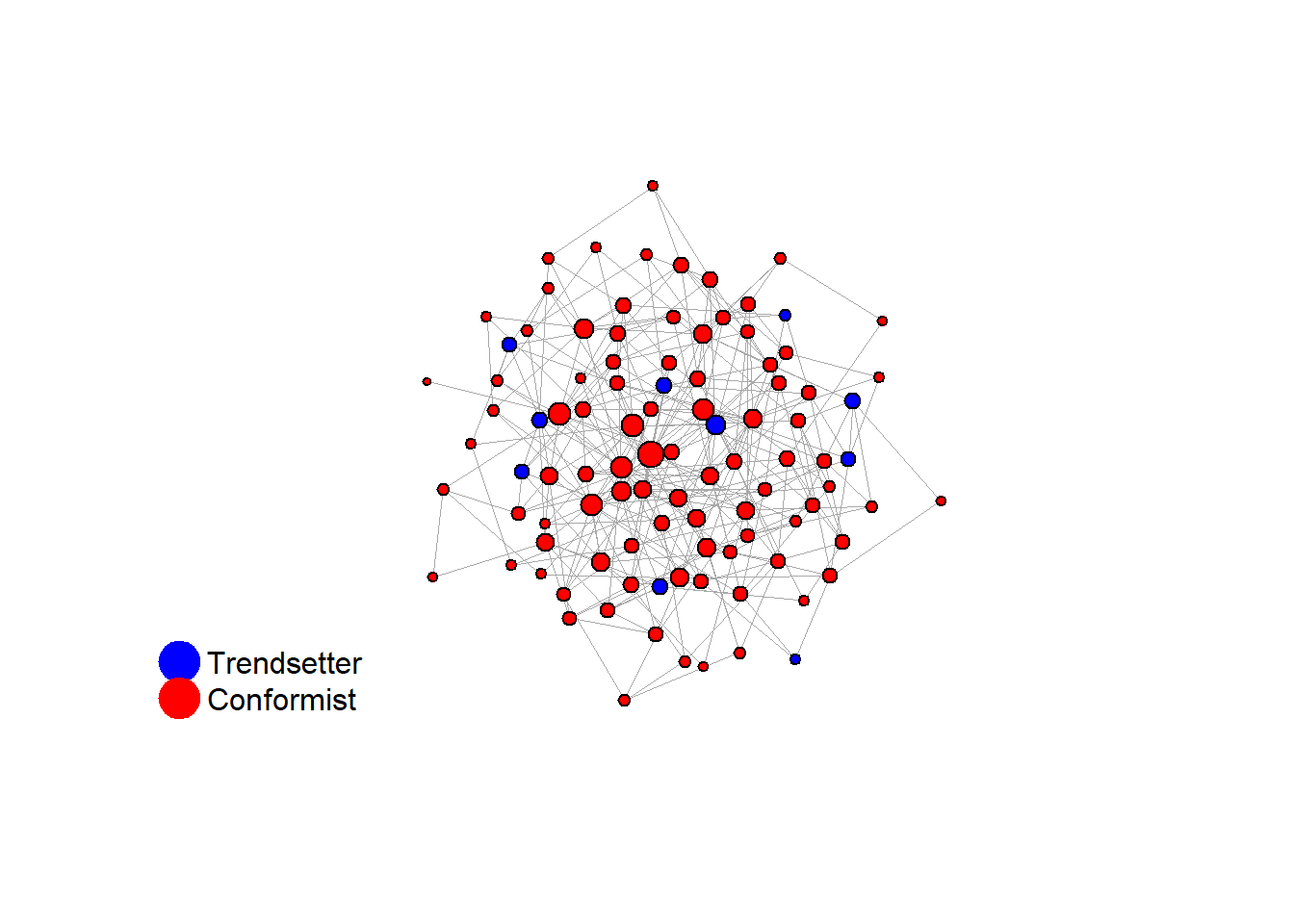
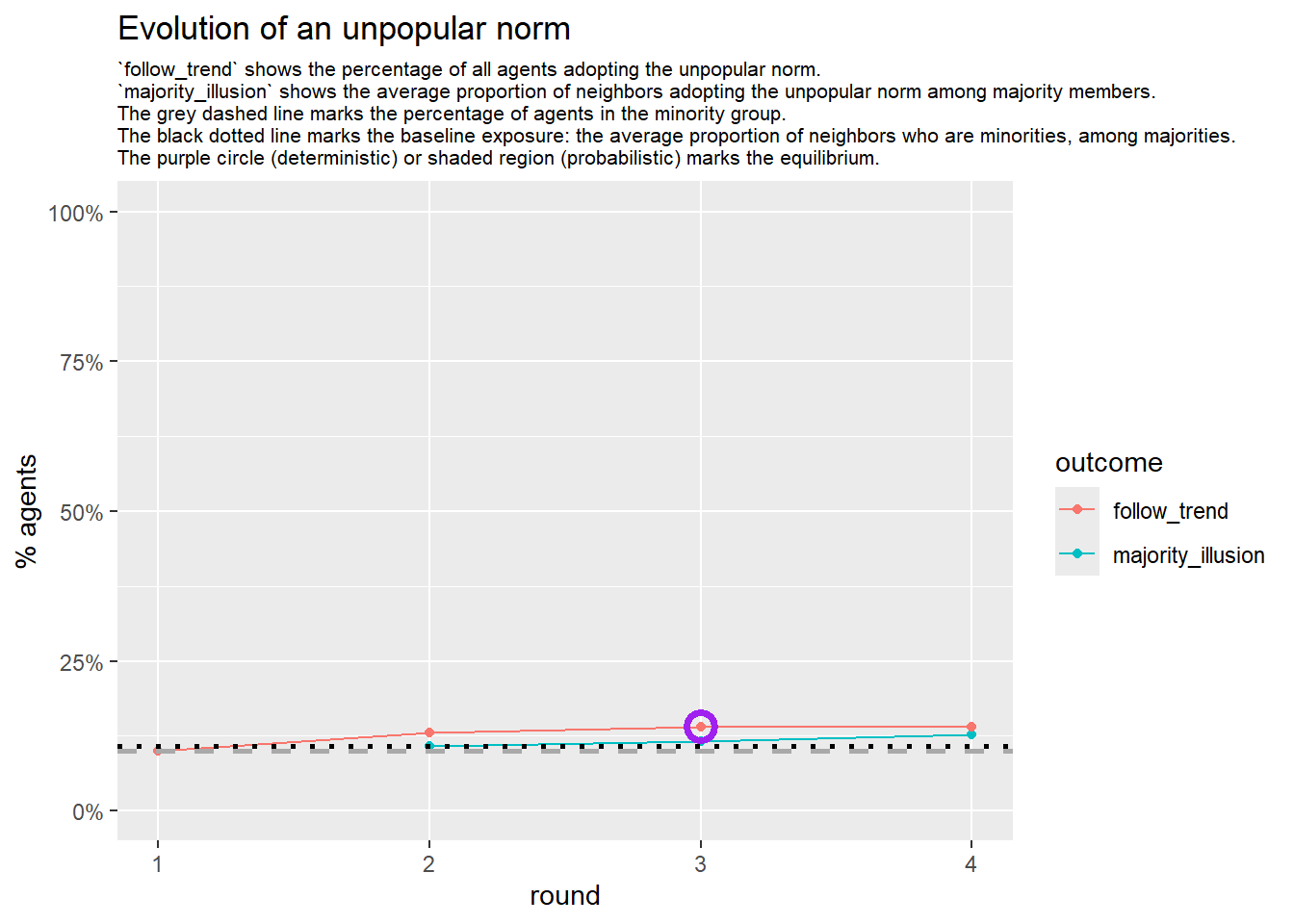
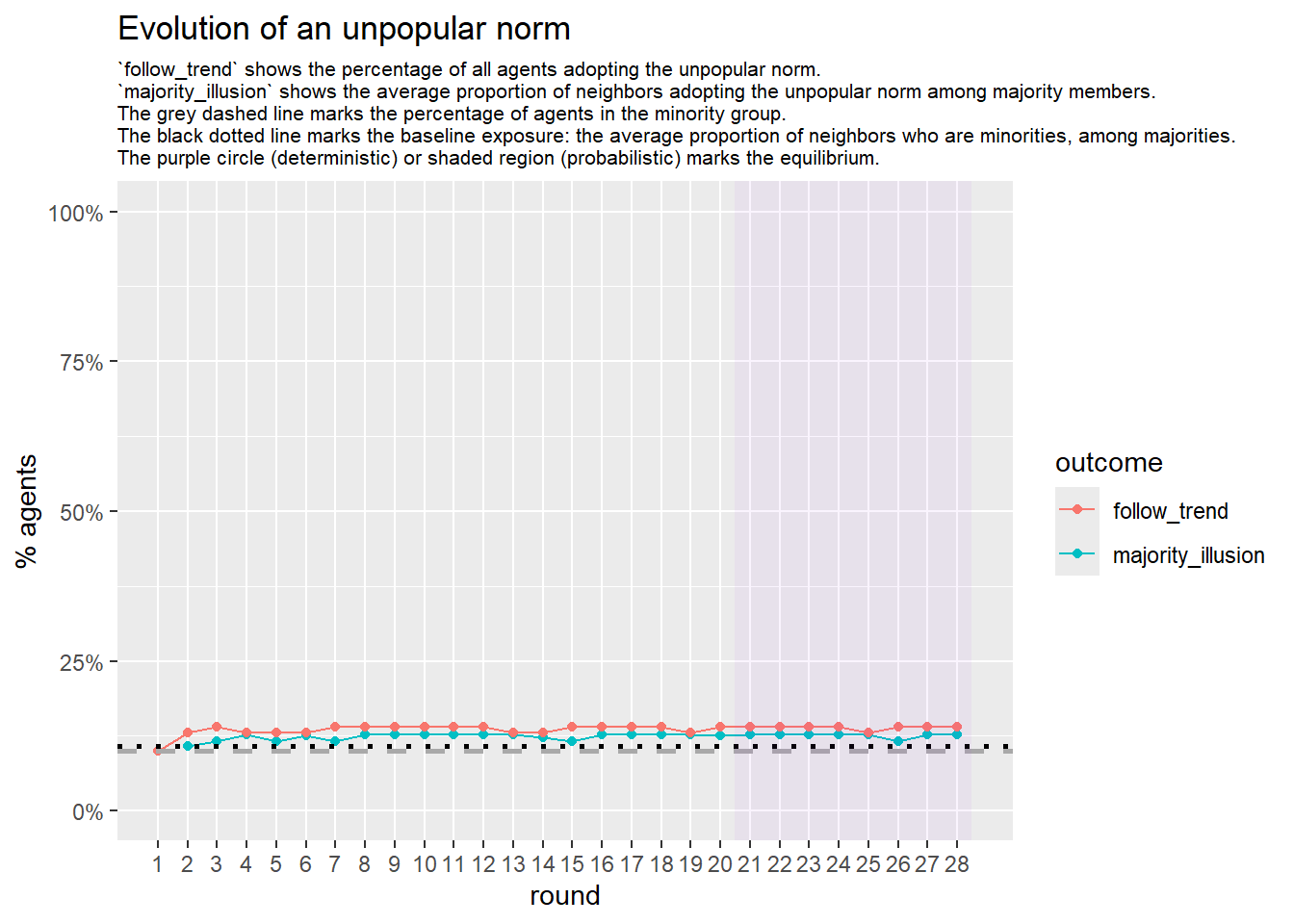
base_seed = 1253281
seed = base_seed + 15
# let's check the distribution of distances to 'trendsetters', among 'conformists'
g <- test$network
trend <- which(V(g)$role == "trendsetter")
conf <- which(V(g)$role == "conformist")
# distances: rows = sources (trendsetters), cols = all vertices
D <- distances(g, v = trend, to = V(g), mode = "all")
Dc <- D[, conf] #keep just conformists
# calculate (a) the distance to the nearest trendsetter and (b) the average distance to
# trendsetters, across conformists
dists <- data.frame(shortest = apply(Dc, 2, min), average = apply(Dc, 2, function(x) mean(x[is.finite(x)])))
dists_long <- pivot_longer(dists, cols = c(shortest, average), names_to = "type", values_to = "distance_to_seed")
# plot the distribution
ggplot(dists_long, aes(x = distance_to_seed, fill = type)) + geom_histogram(bins = 20, alpha = 0.5, position = "identity") +
theme_minimal()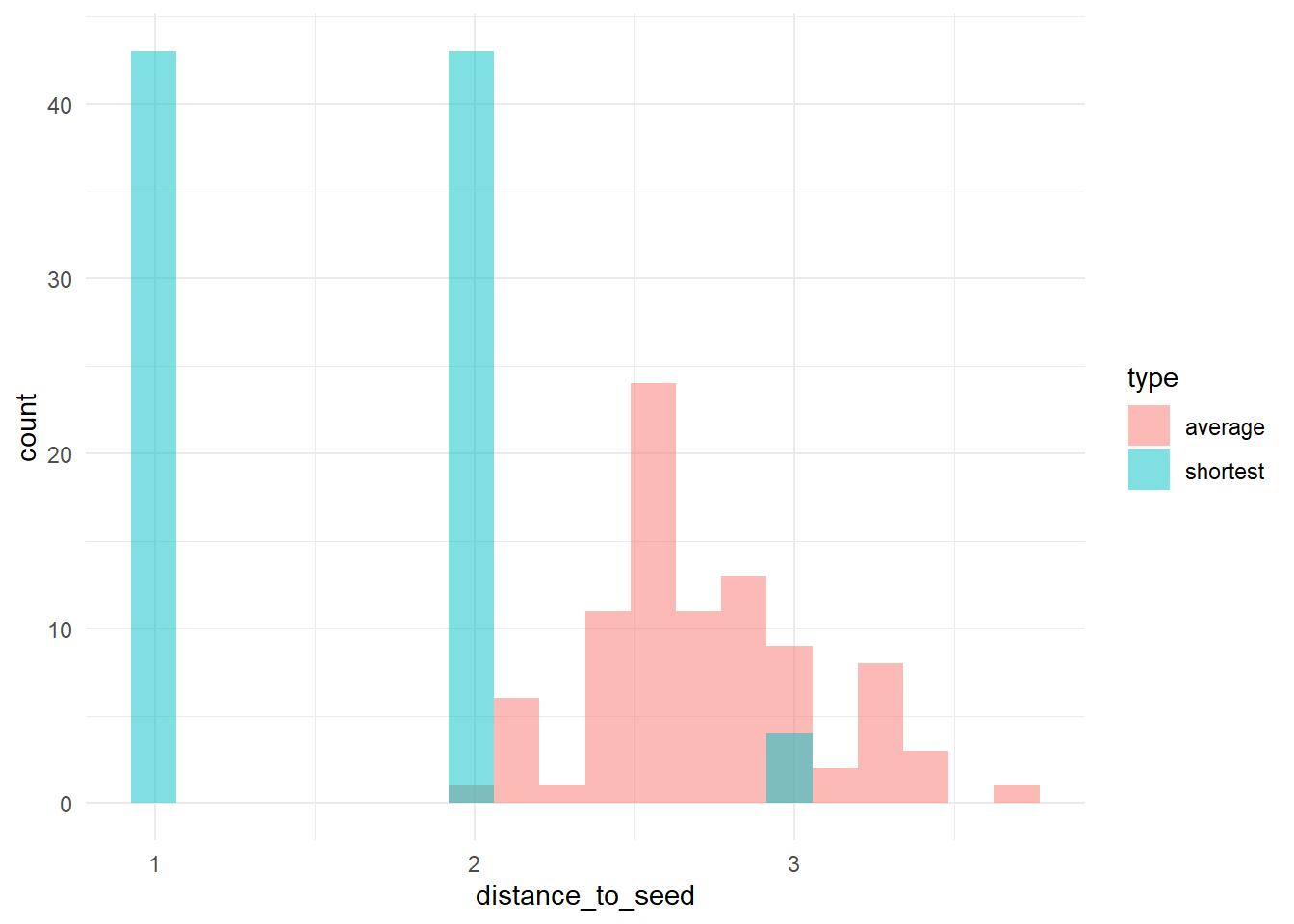
test$stats#> $run
#> [1] 15
#>
#> $seed
#> [1] 1253296
#>
#> $num_nodes
#> [1] 100
#>
#> $num_edges
#> [1] 300
#>
#> $avg_degree
#> [1] 6
#>
#> $sd_degree
#> [1] 2.506557
#>
#> $net_density
#> [1] 0.06060606
#>
#> $net_diameter
#> [1] 5
#>
#> $avg_path_len
#> [1] 2.731919
#>
#> $clust_coeff
#> [1] 0.05963556
#>
#> $assort_deg
#> [1] -0.00588506
#>
#> $deg_trait_cor
#> [1] 0.04009635
#>
#> $components
#> [1] 1table(degree(test$network))#>
#> 1 2 3 4 5 6 7 8 9 10 11 12 15
#> 1 4 11 15 9 23 16 7 5 3 3 2 1# identify trendsetters
ids <- which(V(test$network)$role == "trendsetter")
# check their centrality
sort(as.numeric(cbind(degree(test$network), V(test$network)$role)[ids]))#> [1] 3 4 6 6 6 7 7 7 7 10knitr::include_graphics(paste0("./figures/animation_network_", seed, ".gif"))
# use this as the network structure for an otree session:
# cbind(degree(test$network),V(test$network)$role)
# convert to adjacency matrix
adj_matrix <- as.matrix(as_adjacency_matrix(test$network))
# get roles
role_vector <- ifelse(V(test$network)$role == "trendsetter", 1, 0)
# create a list to store the network data
net <- list(adj_matrix = adj_matrix, role_vector = role_vector)
# save the list as a JSON file
write_json(net, "network_test_n100_random.json")test <- run_one_sw(i = 15, model = "caveman", clique_size = 5, pmin = 0.15, return_network = TRUE)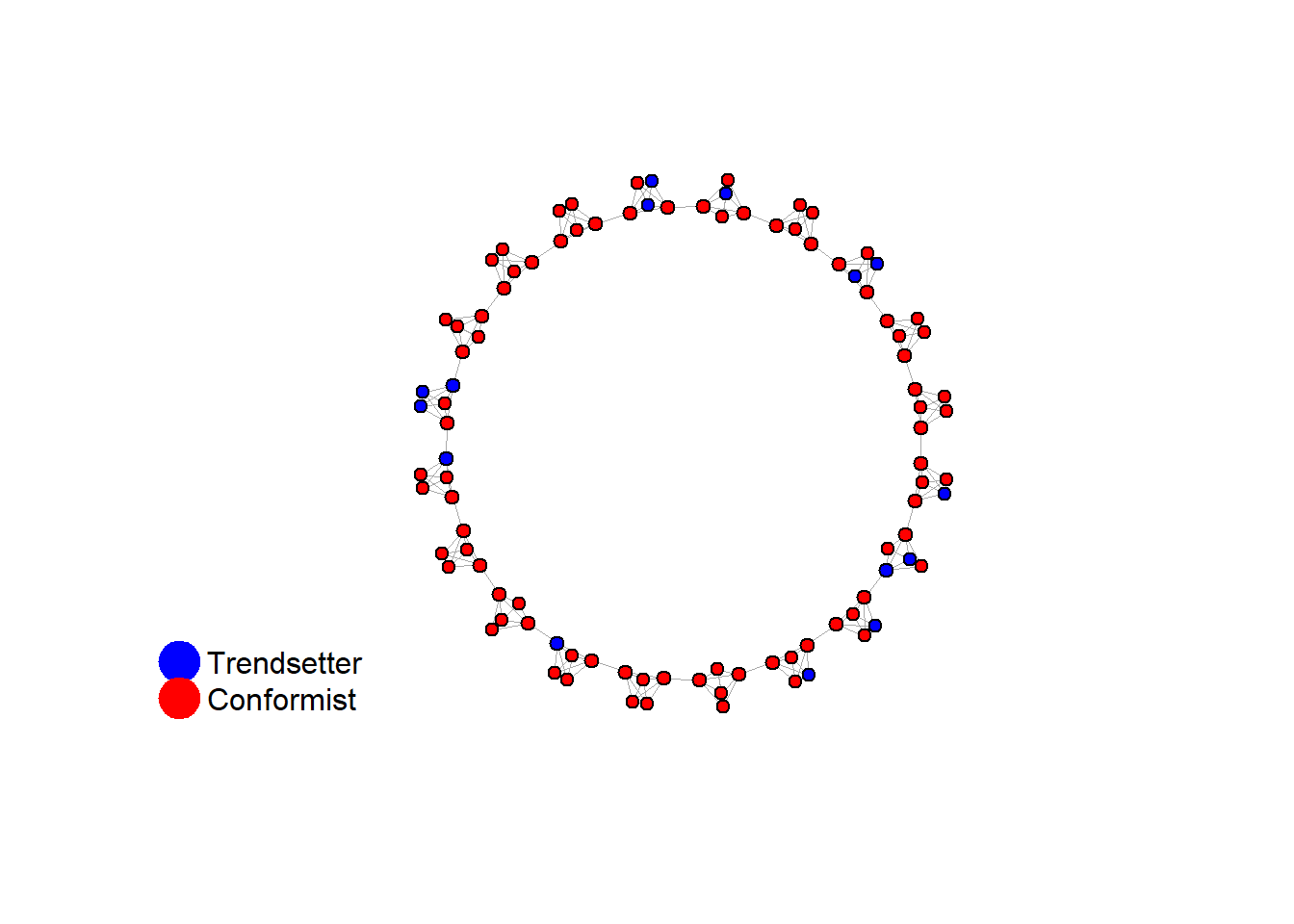
seed = seed + 1
# knitr::include_graphics(paste0('./figures/animation_network_', seed ,'.gif'))LS0tDQp0aXRsZTogIkV4cGVyaW1lbnQiDQpiaWJsaW9ncmFwaHk6IHJlZmVyZW5jZXMuYmliDQpsaW5rLWNpdGF0aW9uczogdHJ1ZQ0KZGF0ZTogIkxhc3QgY29tcGlsZWQgb24gYHIgZm9ybWF0KFN5cy50aW1lKCksICclZC0lbS0lWScpYCINCm91dHB1dDogDQogIGh0bWxfZG9jdW1lbnQ6DQogICAgc2VsZl9jb250YWluZWQ6IHRydWUNCiAgICBjc3M6IHR3ZWFrcy5jc3MNCiAgICB0b2M6IHRydWUNCiAgICB0b2NfZmxvYXQ6IHRydWUNCiAgICBudW1iZXJfc2VjdGlvbnM6IHRydWUNCiAgICB0b2NfZGVwdGg6IDQNCiAgICBjb2RlX2ZvbGRpbmc6IHNob3cNCiAgICBjb2RlX2Rvd25sb2FkOiB5ZXMNCi0tLQ0KDQpgYGB7ciwgZ2xvYmFsc2V0dGluZ3MsIGVjaG89RkFMU0UsIHdhcm5pbmc9RkFMU0UsIHJlc3VsdHM9J2hpZGUnLCBtZXNzYWdlPUZBTFNFfQ0KbGlicmFyeShrbml0cikNCmxpYnJhcnkodGlkeXZlcnNlKQ0Ka25pdHI6Om9wdHNfY2h1bmskc2V0KGVjaG8gPSBUUlVFKQ0Kb3B0c19jaHVuayRzZXQodGlkeS5vcHRzPWxpc3Qod2lkdGguY3V0b2ZmPTEwMCksdGlkeT1UUlVFLCB3YXJuaW5nID0gRkFMU0UsIG1lc3NhZ2UgPSBGQUxTRSxjb21tZW50ID0gIiM+IiwgY2FjaGU9VFJVRSwgY2xhc3Muc291cmNlPWMoInRlc3QiKSwgY2xhc3Mub3V0cHV0PWMoInRlc3QzIikpDQpvcHRpb25zKHdpZHRoID0gMTAwKQ0KcmdsOjpzZXR1cEtuaXRyKCkNCg0KY29sb3JpemUgPC0gZnVuY3Rpb24oeCwgY29sb3IpIHtzcHJpbnRmKCI8c3BhbiBzdHlsZT0nY29sb3I6ICVzOyc+JXM8L3NwYW4+IiwgY29sb3IsIHgpIH0NCmBgYA0KDQpgYGB7ciBrbGlwcHksIGVjaG89RkFMU0UsIGluY2x1ZGU9VFJVRX0NCmtsaXBweTo6a2xpcHB5KHBvc2l0aW9uID0gYygndG9wJywgJ3JpZ2h0JykpDQoja2xpcHB5OjprbGlwcHkoY29sb3IgPSAnZGFya3JlZCcpDQoja2xpcHB5OjprbGlwcHkodG9vbHRpcF9tZXNzYWdlID0gJ0NsaWNrIHRvIGNvcHknLCB0b29sdGlwX3N1Y2Nlc3MgPSAnRG9uZScpDQpgYGANCg0KLS0tDQoNCiMgR2V0dGluZyBzdGFydGVkDQoNClRvIGNvcHkgdGhlIGNvZGUsIGNsaWNrIHRoZSBidXR0b24gaW4gdGhlIHVwcGVyIHJpZ2h0IGNvcm5lciBvZiB0aGUgY29kZS1jaHVua3MuDQoNCiMjIGNsZWFuIHVwDQoNCmBgYHtyLCBjbGVhbl91cCwgcmVzdWx0cz0naGlkZSd9DQpybShsaXN0PWxzKCkpDQpnYygpDQpgYGANCg0KPGJyPg0KDQojIyBjdXN0b20gZnVuY3Rpb25zDQoNCldlIGRlZmluZWQgYSBudW1iZXIgY3VzdG9tIGZ1bmN0aW9ucywgYXQgYHIgeGZ1bjo6ZW1iZWRfZmlsZSgiLi9jdXN0b21fZnVuY3Rpb25zLlIiKWAuDQoNCmBgYHtyLCBjdXN0b21fZnVuY3Rpb25zfQ0Kc291cmNlKCIuL2N1c3RvbV9mdW5jdGlvbnMuUiIpDQpgYGANCg0KPGJyPg0KDQojIyBuZWNlc3NhcnkgcGFja2FnZXMNCg0KLSBgdGlkeXZlcnNlYDogZGF0YSB3cmFuZ2xpbmcNCi0gYGlncmFwaGA6IGdlbmVyYXRlIGFuZCB2aXN1YWxpemUgZ3JhcGhzDQotIGBwYXJhbGxlbGA6IHBhcmFsbGVsIGNvbXB1dGluZyB0byBzcGVlZCB1cCBzaW11bGF0aW9uDQotIGBmb3JlYWNoYDogbG9vcGluZyBpbiBwYXJhbGxlbA0KLSBgZG9QYXJhbGxlbGA6IHBhcmFsbGVsIGJhY2tlbmQgZm9yIGBmb3JlYWNoYA0KLSBgZ2dwbG90MmA6IGRhdGEgdmlzdWFsaXphdGlvbg0KLSBgZ2doNHhgOiBoYWNrcyBmb3IgYGdncGxvdDJgDQotIGBnZ3B1YnJgOiBtYWtlIHZpc3VhbGl6YXRpb25zIHB1YmxpY2F0aW9uLXJlYWR5DQoNCmBgYHtyLCBwYWNrYWdlc30NCnBhY2thZ2VzID0gYygidGlkeXZlcnNlIiwgImlncmFwaCIsICJnZ3Bsb3QyIiwgInBhcmFsbGVsIiwgImRvUGFyYWxsZWwiLCAiZm9yZWFjaCIsICJnZ2g0eCIsICJnZ3B1YnIiLCAicGxvdGx5IiwgIlJDb2xvckJyZXdlciIsICJncmlkIiwgImdyaWRFeHRyYSIsICJwYXRjaHdvcmsiLCAiZ2dwbG90aWZ5IiwgImdncmFwaCIsICJnZ2FuaW1hdGUiLCAiUkNvbG9yQnJld2VyIiwNCiAgICAiZ2d0ZXh0IiwgIm1hZ2ljayIsICJqc29ubGl0ZSIpDQoNCmludmlzaWJsZShmcGFja2FnZS5jaGVjayhwYWNrYWdlcykpDQpybShwYWNrYWdlcykNCmBgYA0KDQotLS0NCg0KIyBFeHBlcmltZW50YWwgY29uZGl0aW9ucw0KDQojIyAxLiBoZWF2eS10YWlsZWQgbmV0d29yayB3aXRoIGNlbnRyYWwgbWlub3JpdGllcw0KDQpMZXQncyB0YWtlIGEgbGlrZWx5LWNhc2UgZm9yIGFuIHVucG9wdWxhciBub3JtLCBnZW5lcmF0ZSBtdWx0aXBsZSBuZXR3b3JrcyBvZiB0aGlzIHRhcmdldCwgYW5kIHNpbXVsYXRlIHRoZSBlbWVyZ2VuY2Ugb2YgdGhlIG5vcm0gZm9sbG93aW5nIGEgZGV0ZXJtaW5pc3RpYyBhbmQgcHJvYmFiaWxpc3RpYyBjaG9pY2UgbW9kZWw6DQoNCmBgYHtyLCBlY2hvPVRSVUUsIGZpZy5zaG93PSdob2xkJywgZmlnLmtlZXA9J2FsbCcsIG1lc3NhZ2U9RkFMU0UsIGZpZy5oZWlnaHQ9NX0NCiMgcGljayBvbmUgY29uZmlndXJhdGlvbiB0aGF0IGxpa2VseSBsZWFkcyB0byBhbiB1bnBvcHVsYXIgbm9ybSwgYW5kIGV4cGxvcmUgbXVsdGlwbGUgJ3NlZWRzJzoNCnJ1bl9vbmVfc2VlZCA8LSBmdW5jdGlvbigNCiAgaSwNCiAgYmFzZV9zZWVkID0gMTI1MzI4MSwNCiAgcGFyYW1zID0gbGlzdChzID0gMTUsIGUgPSAxMCwgdyA9IDQwLCB6ID0gNTAsIGxhbWJkYTEgPSA1LCBsYW1iZGEyID0gMS44KSwNCiAgDQogICMgdHdlYWsgbmV0d29yaw0KICBrX21pbiA9IDIsIA0KICBrX21heCA9IDIwLA0KICBhbHBoYSA9IDIuNCwNCiAgcmhvID0gMC40LA0KICByID0gLTAuMSwNCiAgDQogICMgcmV0cmlldmUgbmV0d29yaw0KICByZXR1cm5fbmV0d29yayA9IEZBTFNFDQogIA0KKSB7DQogICMgZGVyaXZlZCBzZWVkIGZvciB0aGlzIHJ1bg0KICBzZWVkX2kgPC0gYmFzZV9zZWVkICsgaQ0KICBzZXQuc2VlZChzZWVkX2kpDQoNCiAgIyAtLS0gbmV0d29yayBjcmVhdGlvbiAtLS0NCiAgZGVnc2VxIDwtIGZkZWdzZXEoDQogICAgbiAgICAgID0gMTAwLA0KICAgIGFscGhhICA9IGFscGhhLA0KICAgIGtfbWluICA9IGtfbWluLA0KICAgIGtfbWF4ICA9IGtfbWF4LA0KICAgIGRpc3QgICA9ICJsb2ctbm9ybWFsIiwgI3VzZSBsb2ctbm9ybWFsIA0KICAgIHNlZWQgICA9IHNlZWRfaQ0KICApDQoNCiAgbmV0d29yayA8LSBzYW1wbGVfZGVnc2VxKGRlZ3NlcSwgbWV0aG9kID0gInZsIikNCiAgDQogIFYobmV0d29yaykkcm9sZSA8LSBzYW1wbGUoDQogICAgYyhyZXAoInRyZW5kc2V0dGVyIiwgMTApLCByZXAoImNvbmZvcm1pc3QiLCA5MCkpDQogICkNCiAgDQoNCiAgcmV3aXJlZF9uZXR3b3JrIDwtIGZyZXdpcmVfcihuZXR3b3JrLCByLCB2ZXJib3NlID0gRkFMU0UsIG1heF9pdGVyID0gMWU1KQ0KICBmaW5hbF9uZXR3b3JrICAgPC0gZnN3YXBfcmhvKHJld2lyZWRfbmV0d29yaywgcmhvLCB2ZXJib3NlID0gRkFMU0UsIG1heF9pdGVyID0gMWU0KQ0KICANCiAgIyAtLS0gc3RhdHMgLS0tDQogIHN0YXRzIDwtIGxpc3QoDQogICAgcnVuICAgICAgICAgICA9IGksDQogICAgc2VlZCAgICAgICAgICA9IHNlZWRfaSwNCiAgICBudW1fbm9kZXMgICAgID0gdmNvdW50KGZpbmFsX25ldHdvcmspLA0KICAgIG51bV9lZGdlcyAgICAgPSBlY291bnQoZmluYWxfbmV0d29yayksDQogICAgYXZnX2RlZ3JlZSAgICA9IG1lYW4oZGVncmVlKGZpbmFsX25ldHdvcmspKSwNCiAgICBzZF9kZWdyZWUgICAgID0gc2QoZGVncmVlKGZpbmFsX25ldHdvcmspKSwNCiAgICBuZXRfZGVuc2l0eSAgID0gZWRnZV9kZW5zaXR5KGZpbmFsX25ldHdvcmspLA0KICAgIG5ldF9kaWFtZXRlciAgPSBkaWFtZXRlcihmaW5hbF9uZXR3b3JrLCBkaXJlY3RlZCA9IEZBTFNFLCB1bmNvbm5lY3RlZCA9IFRSVUUpLA0KICAgIGF2Z19wYXRoX2xlbiAgPSBhdmVyYWdlLnBhdGgubGVuZ3RoKGZpbmFsX25ldHdvcmssIGRpcmVjdGVkID0gRkFMU0UpLA0KICAgIGNsdXN0X2NvZWZmICAgPSB0cmFuc2l0aXZpdHkoZmluYWxfbmV0d29yaywgdHlwZSA9ICJnbG9iYWwiKSwNCiAgICBhc3NvcnRfZGVnICAgID0gYXNzb3J0YXRpdml0eV9kZWdyZWUoZmluYWxfbmV0d29yayksDQogICAgZGVnX3RyYWl0X2NvciA9IGZkZWd0cmFpdGNvcihmaW5hbF9uZXR3b3JrKSRjb3IsDQogICAgY29tcG9uZW50cyAgICA9IGNvbXBvbmVudHMoZmluYWxfbmV0d29yaykkbm8NCiAgKQ0KICANCiAgZnBsb3RfZ3JhcGgoZmluYWxfbmV0d29yaywgbGF5b3V0ID0gbGF5b3V0X3dpdGhfZnIoZmluYWxfbmV0d29yaykpIA0KICANCiAgDQogICMgLS0tIGluaXRpYWwgYWN0aW9ucyAtLS0NCiAgVihmaW5hbF9uZXR3b3JrKSRhY3Rpb24gPC0gaWZlbHNlKFYoZmluYWxfbmV0d29yaykkcm9sZSA9PSAidHJlbmRzZXR0ZXIiLCAxLCAwKQ0KICANCiAgIyAtLS0gZGV0ZXJtaW5pc3RpYyBzaW11bGF0aW9uIC0tLQ0KICBzaW1fZGV0IDwtIGZhYm0oDQogICAgbmV0d29yayAgICAgID0gZmluYWxfbmV0d29yaywNCiAgICBwYXJhbXMgICAgICAgPSBwYXJhbXMsDQogICAgbWF4X3JvdW5kcyAgID0gNTAsDQogICAgbWlfdGhyZXNob2xkID0gMC40OSwNCiAgICBjaG9pY2VfcnVsZSAgPSAiZGV0ZXJtaW5pc3RpYyIsDQogICAgcGxvdCAgICAgICAgID0gVFJVRSwNCiAgICBoaXN0b3JpZXMgICAgPSBUUlVFDQogICkNCiAgDQogICMgZ2VuZXJhdGUgdGhlIGdpZiBmb3IgdGhlIGN1cnJlbnQgbmV0d29yaw0KICBnaWZfZmlsZW5hbWUgPC0gcGFzdGUwKCIuL2ZpZ3VyZXMvYW5pbWF0aW9uX25ldHdvcmtfIiwgc2VlZF9pLCAiLmdpZiIpDQogIGdpZl9wYXRoIDwtIGZuZXR3b3JrZ2lmKGZpbmFsX25ldHdvcmssIHNpbV9kZXQkZGVjaXNpb25faGlzdG9yeSwgcm91bmRzID0gc2ltX2RldCRlcXVpbGlicml1bSRyb3VuZCwgb3V0cHV0X2RpciA9ICIuL2ZpZ3VyZXMiKQ0KICAjIHJlbmFtZSB0aGUgZ2lmIHRvIG1hdGNoIHRoZSBuYW1pbmcgcGF0dGVybg0KICBmaWxlLnJlbmFtZShnaWZfcGF0aCwgZ2lmX2ZpbGVuYW1lKQ0KDQogIGlmICghaXMubnVsbChzaW1fZGV0JHBsb3QpKSB7DQogICAgcHJpbnQoc2ltX2RldCRwbG90KQ0KICB9DQogIA0KICAjIC0tLSBwcm9iYWJpbGlzdGljIHNpbXVsYXRpb24gLS0tDQogIHNpbV9wcm9iIDwtIGZhYm0oDQogICAgbmV0d29yayAgICAgICAgICAgICAgICA9IGZpbmFsX25ldHdvcmssDQogICAgcGFyYW1zICAgICAgICAgICAgICAgICA9IHBhcmFtcywNCiAgICBtYXhfcm91bmRzICAgICAgICAgICAgID0gMTAwLA0KICAgIG1pX3RocmVzaG9sZCAgICAgICAgICAgPSAwLjQ5LA0KICAgIGNob2ljZV9ydWxlICAgICAgICAgICAgPSAicHJvYmFiaWxpc3RpYyIsDQogICAgc3RhYmxlX3dpbmRvdyAgICAgICAgICA9IDgsICAgIyB0aGUgbGVuZ3RoIG9mIHRoZSB3aW5kb3cgb2YgYWRvcHRpb24gdmFsdWVzDQogICAgcmVxdWlyZWRfc3RhYmxlX3JvdW5kcyA9IDIwLCAjIG51bWJlciBvZiB3aW5kb3dzIG5lZWRlZCB0byBkZWNsYXJlIGVxdWlsaWJyaXVtDQogICAgcGxvdCAgICAgICAgICAgICAgICAgICA9IFRSVUUNCiAgKQ0KICBpZiAoIWlzLm51bGwoc2ltX3Byb2IkcGxvdCkpIHsNCiAgICBwcmludChzaW1fcHJvYiRwbG90KQ0KICB9DQogIA0KICAgcmVzdWx0IDwtIGxpc3QoDQogICAgc2VncmVnYXRpb25fZGV0ICA9IHNpbV9kZXQkZXF1aWxpYnJpdW0kc2VncmVnYXRpb24sDQogICAgc2VncmVnYXRpb25fcHJvYiA9IHNpbV9wcm9iJGVxdWlsaWJyaXVtJHNlZ3JlZ2F0aW9uLA0KICAgIHN0YXRzICAgICAgICAgICAgPSBzdGF0cw0KICApDQogIA0KICBpZiAocmV0dXJuX25ldHdvcmspIHsNCiAgICByZXN1bHQkbmV0d29yayA8LSBmaW5hbF9uZXR3b3JrDQogIH0NCiAgDQogIHJlc3VsdA0KfQ0KDQpgYGANCg0KDQpgYGB7cn0NCnRlc3QgPC0gcnVuX29uZV9zZWVkKDEsIGtfbWluID0gMiwga19tYXggPSAyMCwgYWxwaGEgPSAyLjQsIHJobyA9IDAuNCwgciA9IC0wLjEsIHJldHVybl9uZXR3b3JrID0gVFJVRSkNCg0KYmFzZSA9IDEyNTMyODENCnNlZWQgPSBiYXNlICsgMQ0KYGBgIA0KDQpgYGB7ciwgb3V0LndpZHRoPSI2MCUifQ0Ka25pdHI6OmluY2x1ZGVfZ3JhcGhpY3MocGFzdGUwKCIuL2ZpZ3VyZXMvYW5pbWF0aW9uX25ldHdvcmtfIiwgc2VlZCAsIi5naWYiKSkNCmBgYA0KDQpgYGB7cn0NCnRlc3QgPC0gcnVuX29uZV9zZWVkKDIzLCBrX21pbiA9IDIsIGtfbWF4ID0gODAsIGFscGhhID0gMi40LCByaG8gPSAwLjQsIHIgPSAtMC4xLCByZXR1cm5fbmV0d29yayA9IFRSVUUpDQpzZWVkID0gc2VlZCArIDIyDQojMjINCg0KIzIzLCA4MCAjdHlwaWNhbCBzLWN1cnZlICh0cmFjdGlvbiBhbW9uZyBzdXNjZXB0aWJsZXMuLi4gc3ByYWVkaW5nIGluIHRoZSBtaWRkbGU7IHRhcmdldGluZyBzdXNjZXB0aWJsZXMgbm90IGRpcmVjbHR5IGV4cG9zZWQuKQ0KDQojbGV0J3MgY2hlY2sgdGhlIGRpc3RyaWJ1dGlvbiBvZiBkaXN0YW5jZXMgdG8gJ3RyZW5kc2V0dGVycycsIGFtb25nICdjb25mb3JtaXN0cycNCmcgPC0gdGVzdCRuZXR3b3JrDQp0cmVuZCA8LSB3aGljaChWKGcpJHJvbGUgPT0gInRyZW5kc2V0dGVyIikNCmNvbmYgIDwtIHdoaWNoKFYoZykkcm9sZSA9PSAiY29uZm9ybWlzdCIpDQoNCiMgZGlzdGFuY2VzOiByb3dzID0gc291cmNlcyAodHJlbmRzZXR0ZXJzKSwgY29scyA9IGFsbCB2ZXJ0aWNlcw0KRCA8LSBkaXN0YW5jZXMoZywgdiA9IHRyZW5kLCB0byA9IFYoZyksIG1vZGUgPSAiYWxsIikNCkRjIDwtIERbLCBjb25mXSAja2VlcCBqdXN0IGNvbmZvcm1pc3RzDQoNCiMgY2FsY3VsYXRlIChhKSB0aGUgZGlzdGFuY2UgdG8gdGhlIG5lYXJlc3QgdHJlbmRzZXR0ZXIgYW5kIChiKSB0aGUgYXZlcmFnZSBkaXN0YW5jZSB0byB0cmVuZHNldHRlcnMsIGFjcm9zcyBjb25mb3JtaXN0cw0KZGlzdHMgPC0gZGF0YS5mcmFtZSgNCiAgc2hvcnRlc3QgPSBhcHBseShEYywgMiwgbWluKSwNCiAgYXZlcmFnZSA9IGFwcGx5KERjLCAyLCBmdW5jdGlvbih4KSBtZWFuKHhbaXMuZmluaXRlKHgpXSkpDQopDQpkaXN0c19sb25nIDwtIHBpdm90X2xvbmdlcigNCiAgZGlzdHMsDQogIGNvbHMgPSBjKHNob3J0ZXN0LCBhdmVyYWdlKSwNCiAgbmFtZXNfdG8gPSAidHlwZSIsDQogIHZhbHVlc190byA9ICJkaXN0YW5jZV90b19zZWVkIg0KKQ0KDQojcGxvdCB0aGUgZGlzdHJpYnV0aW9uDQpnZ3Bsb3QoZGlzdHNfbG9uZywgYWVzKHggPSBkaXN0YW5jZV90b19zZWVkLCBmaWxsID0gdHlwZSkpICsNCiAgZ2VvbV9oaXN0b2dyYW0oDQogICAgYmlucyA9IDIwLA0KICAgIGFscGhhID0gMC41LA0KICAgIHBvc2l0aW9uID0gImlkZW50aXR5Ig0KICApICsNCiAgdGhlbWVfbWluaW1hbCgpDQpgYGANCg0KDQpgYGB7cn0NCnRlc3Qkc3RhdHMNCnRhYmxlKGRlZ3JlZSh0ZXN0JG5ldHdvcmspKQ0KDQojaWRlbnRpZnkgdHJlbmRzZXR0ZXJzDQppZHMgPC0gd2hpY2goVih0ZXN0JG5ldHdvcmspJHJvbGUgPT0gInRyZW5kc2V0dGVyIikNCg0KI2NoZWNrIHRoZWlyIGNlbnRyYWxpdHkNCnNvcnQoYXMubnVtZXJpYyhjYmluZChkZWdyZWUodGVzdCRuZXR3b3JrKSwgVih0ZXN0JG5ldHdvcmspJHJvbGUpW2lkc10pKQ0KYGBgDQoNCmBgYHtyLCBvdXQud2lkdGg9IjYwJSJ9DQprbml0cjo6aW5jbHVkZV9ncmFwaGljcyhwYXN0ZTAoIi4vZmlndXJlcy9hbmltYXRpb25fbmV0d29ya18iLCBzZWVkICwiLmdpZiIpKQ0KYGBgDQoNCmBgYHtyLCBldmFsID0gRkFMU0V9DQojIHVzZSB0aGlzIGFzIHRoZSBuZXR3b3JrIHN0cnVjdHVyZSBmb3IgYW4gb3RyZWUgc2Vzc2lvbjoNCiNjYmluZChkZWdyZWUodGVzdCRuZXR3b3JrKSxWKHRlc3QkbmV0d29yaykkcm9sZSkNCg0KIyBjb252ZXJ0IHRvIGFkamFjZW5jeSBtYXRyaXgNCmFkal9tYXRyaXggPC0gYXMubWF0cml4KGFzX2FkamFjZW5jeV9tYXRyaXgodGVzdCRuZXR3b3JrKSkNCg0KI2dldCByb2xlcw0Kcm9sZV92ZWN0b3IgPC0gaWZlbHNlKFYodGVzdCRuZXR3b3JrKSRyb2xlID09ICJ0cmVuZHNldHRlciIsMSwwKQ0KIyBjcmVhdGUgYSBsaXN0IHRvIHN0b3JlIHRoZSBuZXR3b3JrIGRhdGENCm5ldCA8LSBsaXN0KGFkal9tYXRyaXggPSBhZGpfbWF0cml4LCByb2xlX3ZlY3RvciA9IHJvbGVfdmVjdG9yKQ0KIyBzYXZlIHRoZSBsaXN0IGFzIGEgSlNPTiBmaWxlDQp3cml0ZV9qc29uKG5ldCwgIm5ldHdvcmtfdGVzdF9uMTAwLmpzb24iKQ0KYGBgDQoNCg0KYGBge3J9DQp0ZXN0IDwtIHJ1bl9vbmVfc2VlZCgzLCBrX21pbiA9IDIsIGtfbWF4ID0gMjAsIGFscGhhID0gMi40LCByaG8gPSAwLjQsIHIgPSAtMC4xKQ0Kc2VlZCA9IHNlZWQgLSAyMQ0Kc2VlZCA9IHNlZWQgKyAxDQpgYGANCg0KYGBge3IsIG91dC53aWR0aD0iNjAlIn0NCmtuaXRyOjppbmNsdWRlX2dyYXBoaWNzKHBhc3RlMCgiLi9maWd1cmVzL2FuaW1hdGlvbl9uZXR3b3JrXyIsIHNlZWQgLCIuZ2lmIikpDQpgYGANCg0KYGBge3J9DQp0ZXN0IDwtIHJ1bl9vbmVfc2VlZCg0LCBrX21pbiA9IDIsIGtfbWF4ID0gMjAsIGFscGhhID0gMi40LCByaG8gPSAwLjQsIHIgPSAtMC4xKQ0Kc2VlZCA9IHNlZWQgKyAxDQpgYGANCg0KYGBge3IsIG91dC53aWR0aD0iNjAlIn0NCmtuaXRyOjppbmNsdWRlX2dyYXBoaWNzKHBhc3RlMCgiLi9maWd1cmVzL2FuaW1hdGlvbl9uZXR3b3JrXyIsIHNlZWQgLCIuZ2lmIikpDQpgYGANCg0KYGBge3J9DQp0ZXN0IDwtIHJ1bl9vbmVfc2VlZCg1LCBrX21pbiA9IDIsIGtfbWF4ID0gMjAsIGFscGhhID0gMi40LCByaG8gPSAwLjQsIHIgPSAtMC4xKQ0Kc2VlZCA9IHNlZWQgKyAxDQpgYGANCg0KYGBge3IsIG91dC53aWR0aD0iNjAlIn0NCmtuaXRyOjppbmNsdWRlX2dyYXBoaWNzKHBhc3RlMCgiLi9maWd1cmVzL2FuaW1hdGlvbl9uZXR3b3JrXyIsIHNlZWQgLCIuZ2lmIikpDQpgYGANCg0KYGBge3J9DQp0ZXN0IDwtIHJ1bl9vbmVfc2VlZCg2LCBrX21pbiA9IDIsIGtfbWF4ID0gMjAsIGFscGhhID0gMi40LCByaG8gPSAwLjQsIHIgPSAtMC4xKQ0Kc2VlZCA9IHNlZWQgKyAxDQpgYGANCg0KYGBge3IsIG91dC53aWR0aD0iNjAlIn0NCmtuaXRyOjppbmNsdWRlX2dyYXBoaWNzKHBhc3RlMCgiLi9maWd1cmVzL2FuaW1hdGlvbl9uZXR3b3JrXyIsIHNlZWQgLCIuZ2lmIikpDQpgYGANCg0KYGBge3J9DQp0ZXN0IDwtIHJ1bl9vbmVfc2VlZCg3LCBrX21pbiA9IDIsIGtfbWF4ID0gMjAsIGFscGhhID0gMi40LCByaG8gPSAwLjQsIHIgPSAtMC4xKQ0Kc2VlZCA9IHNlZWQgKyAxDQpgYGANCg0KYGBge3IsIG91dC53aWR0aD0iNjAlIn0NCmtuaXRyOjppbmNsdWRlX2dyYXBoaWNzKHBhc3RlMCgiLi9maWd1cmVzL2FuaW1hdGlvbl9uZXR3b3JrXyIsIHNlZWQgLCIuZ2lmIikpDQpgYGANCg0KSW5jcmVhc2UgdGhlIGRlbnNpdHk6DQoNCmBgYHtyfQ0KdGVzdCA8LSBydW5fb25lX3NlZWQoOCwga19taW4gPSA0LCBrX21heCA9IDk5LCBhbHBoYSA9IDIuMSwgcmhvID0gMC41LCByID0gLTAuNCkNCnNlZWQgPSBzZWVkICsgMQ0KYGBgIA0KDQpgYGB7ciwgb3V0LndpZHRoPSI2MCUifQ0Ka25pdHI6OmluY2x1ZGVfZ3JhcGhpY3MocGFzdGUwKCIuL2ZpZ3VyZXMvYW5pbWF0aW9uX25ldHdvcmtfIiwgc2VlZCAsIi5naWYiKSkNCmBgYA0KDQpgYGB7cn0NCnRlc3QgPC0gcnVuX29uZV9zZWVkKDksIGtfbWluID0gNCwga19tYXggPSA5OSwgYWxwaGEgPSAyLjEsIHJobyA9IDAuNSwgciA9IC0wLjQpDQpzZWVkID0gc2VlZCArIDENCmBgYA0KDQpgYGB7ciwgb3V0LndpZHRoPSI2MCUifQ0Ka25pdHI6OmluY2x1ZGVfZ3JhcGhpY3MocGFzdGUwKCIuL2ZpZ3VyZXMvYW5pbWF0aW9uX25ldHdvcmtfIiwgc2VlZCAsIi5naWYiKSkNCmBgYA0KDQpgYGB7cn0NCnRlc3QgPC0gcnVuX29uZV9zZWVkKDEwLCBrX21pbiA9IDQsIGtfbWF4ID0gOTksIGFscGhhID0gMi4xLCByaG8gPSAwLjUsIHIgPSAtMC40KQ0Kc2VlZCA9IHNlZWQgKyAxDQpgYGANCg0KYGBge3IsIG91dC53aWR0aD0iNjAlIn0NCmtuaXRyOjppbmNsdWRlX2dyYXBoaWNzKHBhc3RlMCgiLi9maWd1cmVzL2FuaW1hdGlvbl9uZXR3b3JrXyIsIHNlZWQgLCIuZ2lmIikpDQpgYGANCg0KYGBge3J9DQp0ZXN0IDwtIHJ1bl9vbmVfc2VlZCgxMSwga19taW4gPSA0LCBrX21heCA9IDk5LCBhbHBoYSA9IDIuMSwgcmhvID0gMC41LCByID0gLTAuNCkNCnNlZWQgPSBzZWVkICsgMQ0KYGBgDQoNCmBgYHtyLCBvdXQud2lkdGg9IjYwJSJ9DQprbml0cjo6aW5jbHVkZV9ncmFwaGljcyhwYXN0ZTAoIi4vZmlndXJlcy9hbmltYXRpb25fbmV0d29ya18iLCBzZWVkICwiLmdpZiIpKQ0KYGBgDQoNCmBgYHtyfQ0KdGVzdCA8LSBydW5fb25lX3NlZWQoMTIsIGtfbWluID0gNCwga19tYXggPSA5OSwgYWxwaGEgPSAyLjEsIHJobyA9IDAuNSwgciA9IC0wLjQpDQpzZWVkID0gc2VlZCArIDENCmBgYA0KDQpgYGB7ciwgb3V0LndpZHRoPSI2MCUifQ0Ka25pdHI6OmluY2x1ZGVfZ3JhcGhpY3MocGFzdGUwKCIuL2ZpZ3VyZXMvYW5pbWF0aW9uX25ldHdvcmtfIiwgc2VlZCAsIi5naWYiKSkNCmBgYA0KDQoNCmBgYHtyfQ0KdGVzdCA8LSBydW5fb25lX3NlZWQoMTMsIGtfbWluID0gNCwga19tYXggPSA5OSwgYWxwaGEgPSAyLjEsIHJobyA9IDAuNSwgciA9IC0wLjQpDQpzZWVkID0gc2VlZCArIDENCmBgYA0KDQpgYGB7ciwgb3V0LndpZHRoPSI2MCUifQ0Ka25pdHI6OmluY2x1ZGVfZ3JhcGhpY3MocGFzdGUwKCIuL2ZpZ3VyZXMvYW5pbWF0aW9uX25ldHdvcmtfIiwgc2VlZCAsIi5naWYiKSkNCmBgYA0KDQoNCmBgYHtyfQ0KdGVzdCA8LSBydW5fb25lX3NlZWQoMTQsIGtfbWluID0gNCwga19tYXggPSA5OSwgYWxwaGEgPSAyLjEsIHJobyA9IDAuNSwgciA9IC0wLjQpDQpzZWVkID0gc2VlZCArIDENCmBgYA0KDQpgYGB7ciwgb3V0LndpZHRoPSI2MCUifQ0Ka25pdHI6OmluY2x1ZGVfZ3JhcGhpY3MocGFzdGUwKCIuL2ZpZ3VyZXMvYW5pbWF0aW9uX25ldHdvcmtfIiwgc2VlZCAsIi5naWYiKSkNCmBgYA0KDQojIyAyLiBzbWFsbC13b3JsZHMNCg0KDQpgYGB7ciwgZWNobz1UUlVFLCBmaWcuc2hvdz0naG9sZCcsIGZpZy5rZWVwPSdhbGwnLCBtZXNzYWdlPUZBTFNFLCBmaWcuaGVpZ2h0PTV9DQpydW5fb25lX3N3IDwtIGZ1bmN0aW9uKA0KICBpLA0KICBiYXNlX3NlZWQgPSAxMjUzMjgxLA0KICBtb2RlbCA9ICJ3YXR0cy1zdHJvZ2F0eiIsDQogIGJldGEgPSAwLA0KICBuZWkgPSAzLA0KICBjbGlxdWVfc2l6ZSA9IDUsDQogIHBtaW4gPSAwLjEsDQogIA0KICBwYXJhbXMgPSBsaXN0KHMgPSAxNSwgZSA9IDEwLCB3ID0gNDAsIHogPSA1MCwgbGFtYmRhMSA9IDUsIGxhbWJkYTIgPSAxLjgpLA0KICANCiAgIyByZXRyaWV2ZSBuZXR3b3JrDQogIHJldHVybl9uZXR3b3JrID0gRkFMU0UNCiAgDQopIHsNCiAgIyBkZXJpdmVkIHNlZWQgZm9yIHRoaXMgcnVuDQogIHNlZWRfaSA8LSBiYXNlX3NlZWQgKyBpDQogIHNldC5zZWVkKHNlZWRfaSkNCg0KICBpZiAobW9kZWwgPT0gIndhdHRzLXN0cm9nYXR6Iikgew0KICAgIG5ldHdvcmsgPC0gc2FtcGxlX3NtYWxsd29ybGQoZGltID0gMSwgc2l6ZSA9IDEwMCwgbmVpID0gMywgcCA9IGJldGEpIH0NCiAgZWxzZSBpZiAobW9kZWwgPT0gImNhdmVtYW4iKSB7DQogICAgbmV0d29yayA8LSBzaW11bGF0ZV9jYXZlbWFuKG4gPSAxMDAsIGNsaXF1ZV9zaXplID0gY2xpcXVlX3NpemUpIA0KICB9DQoNCiAgVihuZXR3b3JrKSRyb2xlIDwtIHNhbXBsZSgNCiAgICBjKHJlcCgidHJlbmRzZXR0ZXIiLCByb3VuZCgxMDAqcG1pbikpLCByZXAoImNvbmZvcm1pc3QiLCAxMDAtcm91bmQoMTAwKnBtaW4pKSkNCiAgKQ0KICANCiAgDQogIGZpbmFsX25ldHdvcmsgPC0gbmV0d29yaw0KDQogIA0KICAjIC0tLSBzdGF0cyAtLS0NCiAgc3RhdHMgPC0gbGlzdCgNCiAgICBydW4gICAgICAgICAgID0gaSwNCiAgICBzZWVkICAgICAgICAgID0gc2VlZF9pLA0KICAgIG51bV9ub2RlcyAgICAgPSB2Y291bnQoZmluYWxfbmV0d29yayksDQogICAgbnVtX2VkZ2VzICAgICA9IGVjb3VudChmaW5hbF9uZXR3b3JrKSwNCiAgICBhdmdfZGVncmVlICAgID0gbWVhbihkZWdyZWUoZmluYWxfbmV0d29yaykpLA0KICAgIHNkX2RlZ3JlZSAgICAgPSBzZChkZWdyZWUoZmluYWxfbmV0d29yaykpLA0KICAgIG5ldF9kZW5zaXR5ICAgPSBlZGdlX2RlbnNpdHkoZmluYWxfbmV0d29yayksDQogICAgbmV0X2RpYW1ldGVyICA9IGRpYW1ldGVyKGZpbmFsX25ldHdvcmssIGRpcmVjdGVkID0gRkFMU0UsIHVuY29ubmVjdGVkID0gVFJVRSksDQogICAgYXZnX3BhdGhfbGVuICA9IGF2ZXJhZ2UucGF0aC5sZW5ndGgoZmluYWxfbmV0d29yaywgZGlyZWN0ZWQgPSBGQUxTRSksDQogICAgY2x1c3RfY29lZmYgICA9IHRyYW5zaXRpdml0eShmaW5hbF9uZXR3b3JrLCB0eXBlID0gImdsb2JhbCIpLA0KICAgIGFzc29ydF9kZWcgICAgPSBhc3NvcnRhdGl2aXR5X2RlZ3JlZShmaW5hbF9uZXR3b3JrKSwNCiAgICBkZWdfdHJhaXRfY29yID0gZmRlZ3RyYWl0Y29yKGZpbmFsX25ldHdvcmspJGNvciwNCiAgICBjb21wb25lbnRzICAgID0gY29tcG9uZW50cyhmaW5hbF9uZXR3b3JrKSRubw0KICApDQogIA0KICBzdGF0c19kZiA8LSBkYXRhLmZyYW1lKA0KICAgIE1ldHJpYyA9IG5hbWVzKHN0YXRzKSwNCiAgICBWYWx1ZSAgPSB1bmxpc3Qoc3RhdHMpLA0KICAgIHJvdy5uYW1lcyA9IE5VTEwNCiAgKQ0KICANCiAgI3ByaW50KHN0YXRzX2RmKQ0KICBmcGxvdF9ncmFwaChmaW5hbF9uZXR3b3JrLCBsYXlvdXQgPSBsYXlvdXQua2FtYWRhLmthd2FpKGZpbmFsX25ldHdvcmspKQ0KICANCg0KICAjIC0tLSBpbml0aWFsIGFjdGlvbnMgLS0tDQogIFYoZmluYWxfbmV0d29yaykkYWN0aW9uIDwtIGlmZWxzZShWKGZpbmFsX25ldHdvcmspJHJvbGUgPT0gInRyZW5kc2V0dGVyIiwgMSwgMCkNCiAgDQogICMgLS0tIGRldGVybWluaXN0aWMgc2ltdWxhdGlvbiAtLS0NCiAgc2ltX2RldCA8LSBmYWJtKA0KICAgIG5ldHdvcmsgICAgICA9IGZpbmFsX25ldHdvcmssDQogICAgcGFyYW1zICAgICAgID0gcGFyYW1zLA0KICAgIG1heF9yb3VuZHMgICA9IDUwLA0KICAgIG1pX3RocmVzaG9sZCA9IDAuNDksDQogICAgY2hvaWNlX3J1bGUgID0gImRldGVybWluaXN0aWMiLA0KICAgIHBsb3QgICAgICAgICA9IFRSVUUsDQogICAgaGlzdG9yaWVzICAgID0gVFJVRQ0KICApDQoNCiAgIyBnZW5lcmF0ZSB0aGUgZ2lmIGZvciB0aGUgY3VycmVudCBuZXR3b3JrDQogIGdpZl9maWxlbmFtZSA8LSBwYXN0ZTAoIi4vZmlndXJlcy9hbmltYXRpb25fbmV0d29ya18iLCBzZWVkX2ksICIuZ2lmIikNCiAgZ2lmX3BhdGggPC0gZm5ldHdvcmtnaWYoZmluYWxfbmV0d29yaywgc2ltX2RldCRkZWNpc2lvbl9oaXN0b3J5LCByb3VuZHMgPSBzaW1fZGV0JGVxdWlsaWJyaXVtJHJvdW5kLCBvdXRwdXRfZGlyID0gIi4vZmlndXJlcyIpDQogICMgcmVuYW1lIHRoZSBnaWYgdG8gbWF0Y2ggdGhlIG5hbWluZyBwYXR0ZXJuDQogIGZpbGUucmVuYW1lKGdpZl9wYXRoLCBnaWZfZmlsZW5hbWUpDQogIA0KICANCiAgaWYgKCFpcy5udWxsKHNpbV9kZXQkcGxvdCkpIHsNCiAgICBwcmludChzaW1fZGV0JHBsb3QpDQogIH0NCiAgDQogICMgLS0tIHByb2JhYmlsaXN0aWMgc2ltdWxhdGlvbiAtLS0NCiAgc2ltX3Byb2IgPC0gZmFibSgNCiAgICBuZXR3b3JrICAgICAgICAgICAgICAgID0gZmluYWxfbmV0d29yaywNCiAgICBwYXJhbXMgICAgICAgICAgICAgICAgID0gcGFyYW1zLA0KICAgIG1heF9yb3VuZHMgICAgICAgICAgICAgPSAxMDAsDQogICAgbWlfdGhyZXNob2xkICAgICAgICAgICA9IDAuNDksDQogICAgY2hvaWNlX3J1bGUgICAgICAgICAgICA9ICJwcm9iYWJpbGlzdGljIiwNCiAgICBzdGFibGVfd2luZG93ICAgICAgICAgID0gOCwgICAjIHRoZSBsZW5ndGggb2YgdGhlIHdpbmRvdyBvZiBhZG9wdGlvbiB2YWx1ZXMNCiAgICByZXF1aXJlZF9zdGFibGVfcm91bmRzID0gMjAsICMgbnVtYmVyIG9mIHdpbmRvd3MgbmVlZGVkIHRvIGRlY2xhcmUgZXF1aWxpYnJpdW0NCiAgICBwbG90ICAgICAgICAgICAgICAgICAgID0gVFJVRQ0KICApDQogIGlmICghaXMubnVsbChzaW1fcHJvYiRwbG90KSkgew0KICAgIHByaW50KHNpbV9wcm9iJHBsb3QpDQogIH0NCiAgDQogIA0KICByZXN1bHQgPC0gbGlzdCgNCiAgICBzZWdyZWdhdGlvbl9kZXQgID0gc2ltX2RldCRlcXVpbGlicml1bSRzZWdyZWdhdGlvbiwNCiAgICBzZWdyZWdhdGlvbl9wcm9iID0gc2ltX3Byb2IkZXF1aWxpYnJpdW0kc2VncmVnYXRpb24sDQogICAgc3RhdHMgICAgICAgICAgICA9IHN0YXRzDQogICkNCiAgDQogIGlmIChyZXR1cm5fbmV0d29yaykgew0KICAgIHJlc3VsdCRuZXR3b3JrIDwtIGZpbmFsX25ldHdvcmsNCiAgfQ0KICANCiAgcmVzdWx0DQp9DQpgYGANCg0KDQpgYGB7cn0NCiNyYW5kb20NCnRlc3QgPC0gcnVuX29uZV9zdyhpPTE1LCBiZXRhPTEsIHBtaW49LjEsIHJldHVybl9uZXR3b3JrID0gVFJVRSApDQoNCmJhc2Vfc2VlZCA9IDEyNTMyODENCnNlZWQgPSBiYXNlX3NlZWQgKyAxNQ0KDQojbGV0J3MgY2hlY2sgdGhlIGRpc3RyaWJ1dGlvbiBvZiBkaXN0YW5jZXMgdG8gJ3RyZW5kc2V0dGVycycsIGFtb25nICdjb25mb3JtaXN0cycNCmcgPC0gdGVzdCRuZXR3b3JrDQp0cmVuZCA8LSB3aGljaChWKGcpJHJvbGUgPT0gInRyZW5kc2V0dGVyIikNCmNvbmYgIDwtIHdoaWNoKFYoZykkcm9sZSA9PSAiY29uZm9ybWlzdCIpDQoNCiMgZGlzdGFuY2VzOiByb3dzID0gc291cmNlcyAodHJlbmRzZXR0ZXJzKSwgY29scyA9IGFsbCB2ZXJ0aWNlcw0KRCA8LSBkaXN0YW5jZXMoZywgdiA9IHRyZW5kLCB0byA9IFYoZyksIG1vZGUgPSAiYWxsIikNCkRjIDwtIERbLCBjb25mXSAja2VlcCBqdXN0IGNvbmZvcm1pc3RzDQoNCiMgY2FsY3VsYXRlIChhKSB0aGUgZGlzdGFuY2UgdG8gdGhlIG5lYXJlc3QgdHJlbmRzZXR0ZXIgYW5kIChiKSB0aGUgYXZlcmFnZSBkaXN0YW5jZSB0byB0cmVuZHNldHRlcnMsIGFjcm9zcyBjb25mb3JtaXN0cw0KZGlzdHMgPC0gZGF0YS5mcmFtZSgNCiAgc2hvcnRlc3QgPSBhcHBseShEYywgMiwgbWluKSwNCiAgYXZlcmFnZSA9IGFwcGx5KERjLCAyLCBmdW5jdGlvbih4KSBtZWFuKHhbaXMuZmluaXRlKHgpXSkpDQopDQpkaXN0c19sb25nIDwtIHBpdm90X2xvbmdlcigNCiAgZGlzdHMsDQogIGNvbHMgPSBjKHNob3J0ZXN0LCBhdmVyYWdlKSwNCiAgbmFtZXNfdG8gPSAidHlwZSIsDQogIHZhbHVlc190byA9ICJkaXN0YW5jZV90b19zZWVkIg0KKQ0KDQojcGxvdCB0aGUgZGlzdHJpYnV0aW9uDQpnZ3Bsb3QoZGlzdHNfbG9uZywgYWVzKHggPSBkaXN0YW5jZV90b19zZWVkLCBmaWxsID0gdHlwZSkpICsNCiAgZ2VvbV9oaXN0b2dyYW0oDQogICAgYmlucyA9IDIwLA0KICAgIGFscGhhID0gMC41LA0KICAgIHBvc2l0aW9uID0gImlkZW50aXR5Ig0KICApICsNCiAgdGhlbWVfbWluaW1hbCgpDQpgYGANCg0KDQpgYGB7ciwgb3V0LndpZHRoPSI2MCUifQ0KdGVzdCRzdGF0cw0KdGFibGUoZGVncmVlKHRlc3QkbmV0d29yaykpDQoNCiNpZGVudGlmeSB0cmVuZHNldHRlcnMNCmlkcyA8LSB3aGljaChWKHRlc3QkbmV0d29yaykkcm9sZSA9PSAidHJlbmRzZXR0ZXIiKQ0KDQojY2hlY2sgdGhlaXIgY2VudHJhbGl0eQ0Kc29ydChhcy5udW1lcmljKGNiaW5kKGRlZ3JlZSh0ZXN0JG5ldHdvcmspLCBWKHRlc3QkbmV0d29yaykkcm9sZSlbaWRzXSkpDQoNCmtuaXRyOjppbmNsdWRlX2dyYXBoaWNzKHBhc3RlMCgiLi9maWd1cmVzL2FuaW1hdGlvbl9uZXR3b3JrXyIsIHNlZWQgLCIuZ2lmIikpDQpgYGAgDQoNCg0KYGBge3IsIGV2YWwgPSBGQUxTRX0NCiMgdXNlIHRoaXMgYXMgdGhlIG5ldHdvcmsgc3RydWN0dXJlIGZvciBhbiBvdHJlZSBzZXNzaW9uOg0KI2NiaW5kKGRlZ3JlZSh0ZXN0JG5ldHdvcmspLFYodGVzdCRuZXR3b3JrKSRyb2xlKQ0KDQojIGNvbnZlcnQgdG8gYWRqYWNlbmN5IG1hdHJpeA0KYWRqX21hdHJpeCA8LSBhcy5tYXRyaXgoYXNfYWRqYWNlbmN5X21hdHJpeCh0ZXN0JG5ldHdvcmspKQ0KDQojZ2V0IHJvbGVzDQpyb2xlX3ZlY3RvciA8LSBpZmVsc2UoVih0ZXN0JG5ldHdvcmspJHJvbGUgPT0gInRyZW5kc2V0dGVyIiwxLDApDQojIGNyZWF0ZSBhIGxpc3QgdG8gc3RvcmUgdGhlIG5ldHdvcmsgZGF0YQ0KbmV0IDwtIGxpc3QoYWRqX21hdHJpeCA9IGFkal9tYXRyaXgsIHJvbGVfdmVjdG9yID0gcm9sZV92ZWN0b3IpDQojIHNhdmUgdGhlIGxpc3QgYXMgYSBKU09OIGZpbGUNCndyaXRlX2pzb24obmV0LCAibmV0d29ya190ZXN0X24xMDBfcmFuZG9tLmpzb24iKQ0KYGBgDQoNCmBgYHtyfQ0KdGVzdCA8LSBydW5fb25lX3N3KGk9MTUsIG1vZGVsID0gImNhdmVtYW4iLCBjbGlxdWVfc2l6ZSA9IDUsIHBtaW4gPSAuMTUsIHJldHVybl9uZXR3b3JrID0gVFJVRSkNCnNlZWQgPSBzZWVkICsgMQ0KDQoja25pdHI6OmluY2x1ZGVfZ3JhcGhpY3MocGFzdGUwKCIuL2ZpZ3VyZXMvYW5pbWF0aW9uX25ldHdvcmtfIiwgc2VlZCAsIi5naWYiKSkNCmBgYA0KDQo=
Copyright © Rob Franken